Antibody data
- Antibody Data
- Antigen structure
- References [11]
- Comments [0]
- Validations
- Western blot [2]
- Immunocytochemistry [1]
- Other assay [11]
Submit
Validation data
Reference
Comment
Report error
- Product number
- MA5-23515 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- AGO2 Monoclonal Antibody
- Antibody type
- Monoclonal
- Antigen
- Recombinant full-length protein
- Reactivity
- Human, Mouse
- Host
- Mouse
- Isotype
- IgG
- Vial size
- 100 µg
- Concentration
- 2.42 mg/mL
- Storage
- -20°C or -80°C if preferred
Submitted references Silencing of circHIPK3 Sensitizes Paclitaxel-Resistant Breast Cancer Cells to Chemotherapy by Regulating HK2 Through Targeting miR-1286.
Silencing of hsa_circ_0009035 Suppresses Cervical Cancer Progression and Enhances Radiosensitivity through MicroRNA 889-3p-Dependent Regulation of HOXB7.
miR‑30a‑5p mitigates autophagy by regulating the Beclin‑1/ATG16 pathway in renal ischemia/reperfusion injury.
Down-regulation of the tumor suppressor miR-34a contributes to head and neck cancer by up-regulating the MET oncogene and modulating tumor immune evasion.
lncRNA DSCAM-AS1 facilitates the progression of endometrial cancer via miR-136-5p.
SARS-CoV-2 expresses a microRNA-like small RNA able to selectively repress host genes.
circ_0084927 promotes cervical carcinogenesis by sponging miR-1179 that suppresses CDK2, a cell cycle-related gene.
Knockdown of SNHG14 Alleviates MPP(+)-Induced Injury in the Cell Model of Parkinson's Disease by Targeting the miR-214-3p/KLF4 Axis.
Up-Regulation of FSTL3, Regulated by lncRNA DSCAM-AS1/miR-122-5p Axis, Promotes Proliferation and Migration of Non-Small Cell Lung Cancer Cells.
LINC01436, regulating miR-585 and FBXO11, is an oncogenic lncRNA in the progression of gastric cancer.
Circ-ASH2L promotes tumor progression by sponging miR-34a to regulate Notch1 in pancreatic ductal adenocarcinoma.
Ni J, Xi X, Xiao S, Xiao X
Cancer management and research 2021;13:5573-5585
Cancer management and research 2021;13:5573-5585
Silencing of hsa_circ_0009035 Suppresses Cervical Cancer Progression and Enhances Radiosensitivity through MicroRNA 889-3p-Dependent Regulation of HOXB7.
Zhao X, Dong W, Luo G, Xie J, Liu J, Yu F
Molecular and cellular biology 2021 May 21;41(6):e0063120
Molecular and cellular biology 2021 May 21;41(6):e0063120
miR‑30a‑5p mitigates autophagy by regulating the Beclin‑1/ATG16 pathway in renal ischemia/reperfusion injury.
Fang Y, Zou L, He W
International journal of molecular medicine 2021 Jul;48(1)
International journal of molecular medicine 2021 Jul;48(1)
Down-regulation of the tumor suppressor miR-34a contributes to head and neck cancer by up-regulating the MET oncogene and modulating tumor immune evasion.
Wu X, Cheng YL, Matthen M, Yoon A, Schwartz GK, Bala S, Taylor AM, Momen-Heravi F
Journal of experimental & clinical cancer research : CR 2021 Feb 17;40(1):70
Journal of experimental & clinical cancer research : CR 2021 Feb 17;40(1):70
lncRNA DSCAM-AS1 facilitates the progression of endometrial cancer via miR-136-5p.
Li L, Chen P, Huang B, Cai P
Oncology letters 2021 Dec;22(6):825
Oncology letters 2021 Dec;22(6):825
SARS-CoV-2 expresses a microRNA-like small RNA able to selectively repress host genes.
Pawlica P, Yario TA, White S, Wang J, Moss WN, Hui P, Vinetz JM, Steitz JA
Proceedings of the National Academy of Sciences of the United States of America 2021 Dec 28;118(52)
Proceedings of the National Academy of Sciences of the United States of America 2021 Dec 28;118(52)
circ_0084927 promotes cervical carcinogenesis by sponging miR-1179 that suppresses CDK2, a cell cycle-related gene.
Qu X, Zhu L, Song L, Liu S
Cancer cell international 2020;20:333
Cancer cell international 2020;20:333
Knockdown of SNHG14 Alleviates MPP(+)-Induced Injury in the Cell Model of Parkinson's Disease by Targeting the miR-214-3p/KLF4 Axis.
Zhou S, Zhang D, Guo J, Zhang J, Chen Y
Frontiers in neuroscience 2020;14:930
Frontiers in neuroscience 2020;14:930
Up-Regulation of FSTL3, Regulated by lncRNA DSCAM-AS1/miR-122-5p Axis, Promotes Proliferation and Migration of Non-Small Cell Lung Cancer Cells.
Gao L, Chen X, Wang Y, Zhang J
OncoTargets and therapy 2020;13:2725-2738
OncoTargets and therapy 2020;13:2725-2738
LINC01436, regulating miR-585 and FBXO11, is an oncogenic lncRNA in the progression of gastric cancer.
Zhang Y, Yang G, He X, Chen S, Zhang F, Fang X
Cell biology international 2020 Mar;44(3):882-893
Cell biology international 2020 Mar;44(3):882-893
Circ-ASH2L promotes tumor progression by sponging miR-34a to regulate Notch1 in pancreatic ductal adenocarcinoma.
Chen Y, Li Z, Zhang M, Wang B, Ye J, Zhang Y, Tang D, Ma D, Jin W, Li X, Wang S
Journal of experimental & clinical cancer research : CR 2019 Nov 12;38(1):466
Journal of experimental & clinical cancer research : CR 2019 Nov 12;38(1):466
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
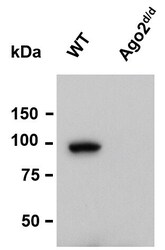
- Experimental details
- Western blot analysis of Ago2 was performed by loading whole cell lysate in 1X LDS sample buffer with 2-ME from 5 x 10^5 mouse transformed pre-B cells. Samples were loaded onto a 4-12 % Bis-Tris polyacrylamide gel (Product # NP0336BOX). Proteins were transferred to nitrocellulose membrane with wet/tank transfer. Membrane was blocked in 5% milk/TBST. The membrane was probe with a monoclonal anti-Ago2 antibody (Product # MA5-23515) at a dilution of 1:1000 in 5% milk/TBST at 4oC overnight, followed by a secondary antibody HRP-anti-mouse at a dilution of 1:10000 at room temperature for 1 hour. Chemiluminescent detection was performed using Pierce™ ECL Western Blotting Substrate (Product # PI-32209). Data courtesy of Thermo Scientific KOL Program.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Whole cell extracts (40 µg) from HeLa cells were analysed by Western blot using the anti-Ago monoclonal antibody antibody (Product # MA5-23515) diluted 1:500 in TBS-Tween containing 5% skimmed milk. The position of the protein of interest is indicated on the right; the marker (in kDa) is shown on the left.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- HeLa cells were stained with the anti-Ago antibody (Product # MA5-23515) and with DAPI. Cells were fixed with 4% formaldehyde for 10’ and blocked with PBS/TX-100 containing 1% BSA. The cells were immunofluorescently labelled with the Ago antibody (middle) diluted 1:500 in blocking solution followed by an anti-mouse antibody conjugated to Alexa594. The left panel shows staining of the nuclei with DAPI. A merge of both stainings is shown on the right.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Coimmunoprecipitation of the miRNA targeted alleles using a AGO2/Argonaute-2 monoclonal antibody (Product # MA5-23515). Briefly, Hela cells were transfected with: pRL-TK-4A + pRL-TK-4G + pcDNA-miR-1 (miR-1 is supposed to target "A" allele); pRL-TK-4A + pRL-TK-4G + pcDNA-miR-1C (miR-1C is supposed to target "G" allele); pRL-TK-4A + pRL-TK-4G + pcDNA-miR-377 (miR-377 is supposed to target neither allele). Cell lysate was immunoprecipitated with Protein G agarose beads incubated with primary antibody. RNA was isolated from the complexes from the input and IP samples, reverse transcribed using random hexamers, and subjected to RT-PCR to confirm enrichment of miRNA targeted allele in IP sample compared to input sample. The PCR products were directly sequenced. The electropherograms at the polymorphic site are shown in the second panel.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Hela cells (10 cm dish per IP) were subjected to UV crosslinking, lysed in 70 µl of 1xPXL buffer (1xPBS, 0.1% SDS, 0.5% Na-deoxycholate, 0.5% NP-40) by incubating on ice for 10 min, then treated with DNase I (ref. Chi SW et al., (2009) Nature 460: 479-486). Dynabeads Protein A (40 µl each) was first incubated with 12 µg anti-mouse IgG as a bridge antibody, briefly washed, and then incubated with a AGO2/Argonaute-2 monoclonal antibody (Product # MA5-23515) (4 or 12 µg), or non-immune mouse IgG control (4 or 12 µg). The cell lysate and Dynabeads were mixed and incubated at 4°C for overnight. The beads were washed twice each at 4°C for 5 min using 500 µl of (i) 1 x PXL buffer, (ii) 5 x PXL (5xPBS, 0.1% SDS, 0.5% Na-deoxycholate, 0.5% NP-40), and (iii) 1 x PNK (50 mM Tris pH7.4, 10 mM MgCl2, 0.5% NP-40). Ten µl of input (10% of starting material), 10 µl of supernatants after immunoprecipitation (10%), and all of immunoprecipitants (~90%) were loaded on SDS-PAGE gel and subjected to Western Blot analysis using Product # MA5-23515 and the ECL plus reagent.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- FIGURE 3 SNHG14 sequestered miR-214-3p via binding to miR-214-3p. (A) Schematic of the miR-214-3p seed sites within SNHG14 identified by the starBase software and mutated the seed sites. (B) Luciferase assays were done in SK-N-SH cells cotransfected with WT-SNHG14 or MUT-SNHG14 and miR-214-3p mimic or miR-NC mimic. (C) RIP assays were performed using anti-Ago2 or anti-IgG antibody in cell lysates of SK-N-SH cells. (D,E) SK-N-SH cells were transfected with si-NC, si-SNHG14, pcDNA, or SNHG14 overexpression vector before MPP + (1 mM, 24 h) stimulation, followed by the measurement of SNHG14 and miR-214-3p levels by qRT-PCR. pcDNA: negative control plasmid, SNHG14: SNHG14 overexpression vector. * P < 0.05.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- FIGURE 5 MiR-214-3p directly interacted with the 3''UTR of KLF4. (A) Schematic of the putative target sequence for miR-214-3p within KLF4 3''UTR and the mutant in the target sequence. (B) Luciferase activities were evaluated in SK-N-SH cells transfected with KLF4 3''UTR-WT or KLF4 3''UTR-MUT together with miR-NC mimic or miR-214-3p mimic. (C) The enrichment levels of miR-214-3p and KLF4 by anti-Ago2 or anti-IgG antibody in cell lysates of SK-N-SK cells. (D,E) KLF4 protein expression was tested by western blot in 6 MPTP-treated mice and 6 normal controls, and primary neuronal cells after treatment with the indicated concentrations of MPP + for 24 h. * P < 0.05.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 3 a-b The in-situ expressions of circ-ASH2L ( a ) and 18S ( b , as a control) in Capan-1 cells. Scale bars = 12.5 mum. c The expressions of indicated miRNAs in indicate treated HEK-293 cells were measured by qRT-PCR. d The prediction for miR-34a binding sites on circ-ASH2L transcript. e Schematic outlining the wild type and mut circ-ASH2L luciferase plasmid. ( f ) Luciferase activity in HEK-293 cells co-transfected with indicated concentration of miR-34a or circ-ASH2L luciferase reporter transcript. Data are showed as the ratio of firefly activity to Renilla luciferase activity. g RNA immunoprecipitation (RIP) experiments were performed using the anti-Ago2 or lgG antibody to immunoprecipitates, the expressions of circ-ASH2L or beta-actin were measured by qRT-PCR. h The schematic diagram of RNA pull-down assay in this study. i circ-ASH2L was pulled down and enriched with miR-34a probe and then detected by qRT-PCR
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 4 miR-34a directly interacts with MET mRNA and downregulates MET. a RNA was isolated from cells, biotin-based pull-down probe for miR-34a-5p or negative control probe at different concentrations to assess interaction of miR-34a-5p with MET in CAL27 cells was used. The enrichment of MET was quantified by qRT-PCR. b and c The miR-34a-5p overexpressing CAL27 cells were subjected to an anti-Ago2 CHIP assay. Levels of MET and miR-34a-5p were quantified by RIP-qPCR. d Dual-luciferase assay was performed in cells transfected with miR-34a-5p mimic or a control miR mimic and a luciferase reporter construct encoding the luciferase gene fused either to a mutated MET 3' UTR (MUT) or a wild-type MET 3' UTR (WT). e and f MET expression and miR-34a-5p expression in HNSCC cell lines were quantified by a TaqMan microRNA assay. g miR-34a-5p mimic was introduced to CAL27 cells by electroporation (25 nM). Levels of miR-34a-5p was quantified by a TaqMan microRNA assay after 12 h. h MET mRNA expression was quantified 24 h after over-expression of miR-34a-5p mimic. i Western blot measuring MET protein expression with and without expression of miR-34a-5p mimic. All experiments were performed in CAL27 cells. All experiments were performed in triplicate. Data represent results of at least three independent experiments. Data is presented as mean +- SD. * indicates p < 0.05
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3 miR-30a-5p directly interacts with the 3'UTR of Beclin-1. (A) The potential binding sites of miR-30a-5p and the 3'UTR of Beclin-1. (B) 293T cells were co-transfected with the pGLO-Beclin-1-WT or pGLO-Beclin-1-Mut plasmid and miR-30a-5p mimic or miR-30a-5p mimic-nc, and luciferase activity was detected (n=3). (C) HK-2 cells were transfected with a miR-30a-5p mimic or miR-30a-5p inhibitor, and the protein levels of Beclin-1 were determined by western blot and semi-quantified. The untransfected cells were used in control group (n=3). (D) mRNA expression levels of Beclin-1 were detected by RT-qPCR. (E) RNA immunoprecipitation assay was performed using anti-Ago2 in HK-2 cells transfected with miR-30a-5p mimic or mimic-nc. miR-30a-5p and 3'UTR of Beclin-1 mRNA expression levels in the anti-Ago2 immunoprecipitated products were measured by RT-qPCR. * P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- FIG 6 hsa_circ_0009035 in CC cells directly targeted miR-889-3p by binding to miR-889-3p. (A) Venn diagram of the putative miRNAs of hsa_circ_0009035 predicted by Circinteractome and Starbase softwares. (B and C) The expression levels of miR-1296, miR-182, miR-653, and miR-889-3p by qRT-PCR in sh-NC-infected or sh-hsa_circ_0009035-transduced HeLa and Siha cells. (D) RNA pulldown assays in HeLa cells using Bio-anti-hsa_circ_0009035 or Bio-NC. (E) Sequence of miR-889-3p and the putative miR-889-3p-binding sequence within hsa_circ_0009035. (F and G) Relative miR-889-3p expression determined by qRT-PCR in HeLa and Siha cells transfected with negative-control plasmid pCD5-ciR, an hsa_circ_0009035-overexpressing plasmid (hsa_circ_0009035), or a mutant hsa_circ_0009035-overexpressing plasmid (MUT-hsa_circ_0009035). (H) RIP assay in HeLa and Siha cells using IgG or AGO2 antibody. (I) RNA pulldown assays in HeLa cells using Bio-NC or Bio-miR-889-3p. (J and K) qRT-PCR analysis of hsa_circ_0009035 expression in cells transfected with miR-NC mimic, miR-889-3p mimic, anti-miR-889-3p, or anti-miR-NC. (L) The level of pre-miR-899 by qRT-PCR in cells transduced with sh-NC, sh-hsa_circ_0009035, or sh-hsa_circ_0009035#2. Relative miR-889-3p expression by qRT-PCR in 82 pairs of CC tissues and adjacent normal tissues (M) and CC tissues from 36 primary patients (defined as radiation-sensitive CC) and 46 recurrent patients after radiation treatment (defined as radiation-resistant CC) (N). (O) Corr
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 4. DSCAM-AS1 specifically binds miR-136-5p and regulates miR-136-5p expression. (A) The putative binding sites between DSCAM-AS1 and miR-136-5p and the mutant sequences of DSCAM-AS1 are presented. (B) miR-136-5p levels were assessed in HEC-1-B and HEC-1-A cells after miR-136-5p mimic or inhibitor transfection. Luciferase activity was detected in (C) HEC-1-B and (D) HEC-1-A cells co-transfected with WT-DSCAM-AS1 or MT-DSCAM-AS1 and miR-136-5p mimic or mi-con. ***P
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- 3 LINC01436 targeted miR-585 in gastric cancer (GC) Cells . (A) Based on data from the StarBase online database, we predicted the binding sites of LINC01436 to miR-585. (B) The expression of miR-585 in paired non-cancerous tissues of GC was detected by quantitative real-time polymerase chain reaction (qRT-PCR). (C) The expression of miR-585 in five GC cell lines and the gastric mucosal epithelial cell was detected by qRT-PCR. (D) After knocking down or overexpressing LINC01436, the expression of miR-585 in BGC823 and AGS cells was detected by qPCR. (E) The coexistence of LINC01436 and miR-585 in GC was identified by the RNA immunoprecipitation assay. (F) Luciferase reporter gene assay confirmed the binding ability between the wild-type binding sites of miR-585 and LINC01436. (G) The expression of LINC01436 and miR-585 was detected by qRT-PCR to verify the correlation of their expression. * P < 0.05, ** P < 0.01, *** P < 0.001.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- 5 LINC01436 as ceRNA regulated FBXO11 expressions by targeting miR-585 . (A) The binding point between miR-585 and FBXO11 was predicted using microRNA.org . (B) Immunohistochemistry was used to detect the expression of FBXO11 in cancer tissues and adjacent normal tissues of gastric cancer (GC) patients. (C) The expression of FBXO11 in GC cells was lower than that in normal gastric mucosa cells by quantitative real-time polymerase chain reaction (qRT-PCR). (D) The correlation between LINC01436 and FBXO11 expression was verified by qRT-PCR. (E) The correlation between the expression of miR-585 and FBXO11 was verified by qRT-PCR. (F, G) RIP measurements showed that LINC01436, miR-585, and FBXO11 coexisted in GC cells. (H) Luciferase reporter gene assay confirmed the binding ability between the wild-type binding sites of miR-585 and FBXO11 . (I) The effect of overexpression of mir-585 or overexpression of LINC01436 on the expression of FBXO11 was detected by qRT-PCR. (J) Overexpression of miR-585 or LINC01436 resulted in down-regulation or up-regulation of FBXO11 protein, respectively. * P < 0.05, ** P < 0.01, *** P < 0.001, n.s P > 0.05.
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot Immunoprecipitation
Immunoprecipitation Immunohistochemistry
Immunohistochemistry