Antibody data
- Antibody Data
- Antigen structure
- References [92]
- Comments [0]
- Validations
- Western blot [1]
- Immunohistochemistry [3]
- Other assay [46]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 13-1600 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- Ubiquitin Monoclonal Antibody (Ubi-1)
- Antibody type
- Monoclonal
- Antigen
- Purifed from natural sources
- Description
- 13-1600 recognizes ubiquitin, both conjugated and unconjugated. It reacts with a single chain 8.5 kDa protein. The ubiquitin molecule appears to be present in all eukaryotic cells and has an identical primary structure in all animals. Ubiquitin is present in the nucleus, cytoplasm, and on cell surface membranes.
- Antibody clone number
- Ubi-1
- Concentration
- 0.5 mg/mL
Submitted references Activation of the ubiquitin-proteasome system contributes to oculopharyngeal muscular dystrophy through muscle atrophy.
Interfering with mitochondrial dynamics sensitizes glioblastoma multiforme to temozolomide chemotherapy.
SPTBN1 inhibits inflammatory responses and hepatocarcinogenesis via the stabilization of SOCS1 and downregulation of p65 in hepatocellular carcinoma.
Suppressing TRAP1 sensitizes glioblastoma multiforme cells to temozolomide.
GMPPA defects cause a neuromuscular disorder with α-dystroglycan hyperglycosylation.
The ubiquitin isopeptidase USP10 deubiquitinates LC3B to increase LC3B levels and autophagic activity.
The Pah-R261Q mouse reveals oxidative stress associated with amyloid-like hepatic aggregation of mutant phenylalanine hydroxylase.
The degradation-promoting roles of deubiquitinases Ubp6 and Ubp3 in cytosolic and ER protein quality control.
Genetic Selection Based on a Ste6(*)C-HA-Ura3 Substrate Identifies New Cytosolic Quality Control Alleles in Saccharomyces cerevisiae.
IL-1β suppression of VE-cadherin transcription underlies sepsis-induced inflammatory lung injury.
Negative regulation of autophagy by UBA6-BIRC6-mediated ubiquitination of LC3.
Replication of multiple system atrophy prions in primary astrocyte cultures from transgenic mice expressing human α-synuclein.
APOPT1/COA8 assists COX assembly and is oppositely regulated by UPS and ROS.
A Mycobacterium tuberculosis surface protein recruits ubiquitin to trigger host xenophagy.
Ubiquitin-specific peptidase 2a (USP2a) deubiquitinates and stabilizes β-catenin.
Cardiac-specific ablation of the E3 ubiquitin ligase Mdm2 leads to oxidative stress, broad mitochondrial deficiency and early death.
Cycloheximide promotes paraptosis induced by inhibition of cyclophilins in glioblastoma multiforme.
Regulation of the ubiquitylation and deubiquitylation of CREB-binding protein modulates histone acetylation and lung inflammation.
Small chaperons and autophagy protected neurons from necrotic cell death.
Fbxl18 targets LRRK2 for proteasomal degradation and attenuates cell toxicity.
Post-translational modifications of Annexin A2 are linked to its association with perinuclear nonpolysomal mRNP complexes.
The E3 ligase Mule protects the heart against oxidative stress and mitochondrial dysfunction through Myc-dependent inactivation of Pgc-1α and Pink1.
Drosophila Mitf regulates the V-ATPase and the lysosomal-autophagic pathway.
Comparison of mHTT Antibodies in Huntington's Disease Mouse Models Reveal Specific Binding Profiles and Steady-State Ubiquitin Levels with Disease Development.
Aberrant Splicing Promotes Proteasomal Degradation of L-type Ca(V)1.2 Calcium Channels by Competitive Binding for Ca(V)β Subunits in Cardiac Hypertrophy.
Multiple myeloma-derived MMP-13 mediates osteoclast fusogenesis and osteolytic disease.
Reactive oxygen species exert opposite effects on Tyr23 phosphorylation of the nuclear and cortical pools of annexin A2.
NQO1 inhibits proteasome-mediated degradation of HIF-1α.
Optineurin deficiency in mice is associated with increased sensitivity to Salmonella but does not affect proinflammatory NF-κB signaling.
Phosphorylation of OPTN by TBK1 enhances its binding to Ub chains and promotes selective autophagy of damaged mitochondria.
mRNA 3'-UTR shortening is a molecular signature of mTORC1 activation.
The cell cycle regulator 14-3-3σ opposes and reverses cancer metabolic reprogramming.
Identification of a subnuclear body involved in sequence-specific cytokine RNA processing.
SLAP displays tumour suppressor functions in colorectal cancer via destabilization of the SRC substrate EPHA2.
CSN6 drives carcinogenesis by positively regulating Myc stability.
Ubiquitin-dependent regulation of MEKK2/3-MEK5-ERK5 signaling module by XIAP and cIAP1.
Monoclonal antibody against the ectodomain of E-cadherin (DECMA-1) suppresses breast carcinogenesis: involvement of the HER/PI3K/Akt/mTOR and IAP pathways.
Drosophila Fip200 is an essential regulator of autophagy that attenuates both growth and aging.
Dissecting the pathways that destabilize mutant p53: the proteasome or autophagy?
An intrinsically disordered region of the acetyltransferase p300 with similarity to prion-like domains plays a role in aggregation.
Mitochondrial division ensures the survival of postmitotic neurons by suppressing oxidative damage.
Analysis of nuclear factor-κB (NF-κB) essential modulator (NEMO) binding to linear and lysine-linked ubiquitin chains and its role in the activation of NF-κB.
Gene expression levels assessed by CA1 pyramidal neuron and regional hippocampal dissections in Alzheimer's disease.
Optineurin is colocalized with ubiquitin in Marinesco bodies.
Dietary downregulation of mutant p53 levels via glucose restriction: mechanisms and implications for tumor therapy.
Subunit 6 of the COP9 signalosome promotes tumorigenesis in mice through stabilization of MDM2 and is upregulated in human cancers.
Synaptic dysbindin-1 reductions in schizophrenia occur in an isoform-specific manner indicating their subsynaptic location.
Protein interactions among Fe65, the low-density lipoprotein receptor-related protein, and the amyloid precursor protein.
A photoconvertible fluorescent reporter to track chaperone-mediated autophagy.
Amyloid precursor-like protein 2 cleavage contributes to neuronal intranuclear inclusions and cytotoxicity in spinocerebellar ataxia-7 (SCA7).
SCA1-like disease in mice expressing wild-type ataxin-1 with a serine to aspartic acid replacement at residue 776.
Neurodegeneration by polyglutamine Atrophin is not rescued by induction of autophagy.
Psh1 is an E3 ubiquitin ligase that targets the centromeric histone variant Cse4.
Different modes of ubiquitination of the adaptor TRAF3 selectively activate the expression of type I interferons and proinflammatory cytokines.
Chemical genetics approach to restoring p27Kip1 reveals novel compounds with antiproliferative activity in prostate cancer cells.
Autophagy in load-induced heart disease.
Interactions of the NPXY microdomains of the low density lipoprotein receptor-related protein 1.
Early-onset and robust amyloid pathology in a new homozygous mouse model of Alzheimer's disease.
Thioredoxin-related Protein 32 is an arsenite-regulated Thiol Reductase of the proteasome 19 S particle.
IKK phosphorylates Huntingtin and targets it for degradation by the proteasome and lysosome.
Regulation of mitochondrial morphology by USP30, a deubiquitinating enzyme present in the mitochondrial outer membrane.
The NER protein Rad33 shows functional homology to human Centrin2 and is involved in modification of Rad4.
Rapid identification of differentiation markers from whole epithelial cells by matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry and statistical analysis.
Geldanamycin induces G2 arrest in U87MG glioblastoma cells through downregulation of Cdc2 and cyclin B1.
PDLIM2-mediated termination of transcription factor NF-kappaB activation by intranuclear sequestration and degradation of the p65 subunit.
Kinase MEKK1 is required for CD40-dependent activation of the kinases Jnk and p38, germinal center formation, B cell proliferation and antibody production.
Allosteric activation of human glucokinase by free polyubiquitin chains and its ubiquitin-dependent cotranslational proteasomal degradation.
RhoB facilitates c-Myc turnover by supporting efficient nuclear accumulation of GSK-3.
Activation of the E3 ubiquitin ligase Itch through a phosphorylation-induced conformational change.
Experimental tests to definitively determine ubiquitylation of a substrate.
Defining the role of ubiquitin-interacting motifs in the polyglutamine disease protein, ataxin-3.
Geldanamycin-induced degradation of Chk1 is mediated by proteasome.
Methylation silencing of SOCS-3 promotes cell growth and migration by enhancing JAK/STAT and FAK signalings in human hepatocellular carcinoma.
Involvement of the p97-Ufd1-Npl4 complex in the regulated endoplasmic reticulum-associated degradation of inositol 1,4,5-trisphosphate receptors.
Role of p97 AAA-ATPase in the retrotranslocation of the cholera toxin A1 chain, a non-ubiquitinated substrate.
Histomathematical analysis of clinical specimens: challenges and progress.
Similar patterns of mitochondrial vulnerability and rescue induced by genetic modification of alpha-synuclein, parkin, and DJ-1 in Caenorhabditis elegans.
Proteasome mediates dopaminergic neuronal degeneration, and its inhibition causes alpha-synuclein inclusions.
Distinct steps in dislocation of luminal endoplasmic reticulum-associated degradation substrates: roles of endoplamic reticulum-bound p97/Cdc48p and proteasome.
A signalling pathway controlling c-Myc degradation that impacts oncogenic transformation of human cells.
Conformational change and mitochondrial translocation of Bax accompany proteasome inhibitor-induced apoptosis of chronic lymphocytic leukemic cells.
Acquired FANCA dysfunction and cytogenetic instability in adult acute myelogenous leukemia.
blue cheese mutations define a novel, conserved gene involved in progressive neural degeneration.
Ubiquitylation of BAG-1 suggests a novel regulatory mechanism during the sorting of chaperone substrates to the proteasome.
Immunohistochemical study of p53-associated proteins in rat brain following lithium-pilocarpine status epilepticus.
Inhibition of Src family kinases blocks epidermal growth factor (EGF)-induced activation of Akt, phosphorylation of c-Cbl, and ubiquitination of the EGF receptor.
Interferon-gamma-mediated activation and ubiquitin-proteasome-dependent degradation of PPARgamma in adipocytes.
Regulation of the ubiquitin-conjugating enzyme hHR6A by CDK-mediated phosphorylation.
Cancer-predisposing mutations within the RING domain of BRCA1: loss of ubiquitin protein ligase activity and protection from radiation hypersensitivity.
Proteasomes regulate erythropoietin receptor and signal transducer and activator of transcription 5 (STAT5) activation. Possible involvement of the ubiquitinated Cis protein.
The Ras-specific exchange factors mouse Sos1 (mSos1) and mSos2 are regulated differently: mSos2 contains ubiquitination signals absent in mSos1.
Identification of a neural alpha-catenin as a key regulator of cadherin function and multicellular organization.
Ribot C, Soler C, Chartier A, Al Hayek S, Naït-Saïdi R, Barbezier N, Coux O, Simonelig M
PLoS genetics 2022 Jan;18(1):e1010015
PLoS genetics 2022 Jan;18(1):e1010015
Interfering with mitochondrial dynamics sensitizes glioblastoma multiforme to temozolomide chemotherapy.
Wang N, Huang R, Yang K, He Y, Gao Y, Dong D
Journal of cellular and molecular medicine 2022 Feb;26(3):893-912
Journal of cellular and molecular medicine 2022 Feb;26(3):893-912
SPTBN1 inhibits inflammatory responses and hepatocarcinogenesis via the stabilization of SOCS1 and downregulation of p65 in hepatocellular carcinoma.
Lin L, Chen S, Wang H, Gao B, Kallakury B, Bhuvaneshwar K, Cahn K, Gusev Y, Wang X, Wu Y, Marshall JL, Zhi X, He AR
Theranostics 2021;11(9):4232-4250
Theranostics 2021;11(9):4232-4250
Suppressing TRAP1 sensitizes glioblastoma multiforme cells to temozolomide.
Wang N, Zhu P, Huang R, Sun L, Dong D, Gao Y
Experimental and therapeutic medicine 2021 Nov;22(5):1246
Experimental and therapeutic medicine 2021 Nov;22(5):1246
GMPPA defects cause a neuromuscular disorder with α-dystroglycan hyperglycosylation.
Franzka P, Henze H, Jung MJ, Schüler SC, Mittag S, Biskup K, Liebmann L, Kentache T, Morales J, Martínez B, Katona I, Herrmann T, Huebner AK, Hennings JC, Groth S, Gresing L, Horstkorte R, Marquardt T, Weis J, Kaether C, Mutchinick OM, Ori A, Huber O, Blanchard V, von Maltzahn J, Hübner CA
The Journal of clinical investigation 2021 May 3;131(9)
The Journal of clinical investigation 2021 May 3;131(9)
The ubiquitin isopeptidase USP10 deubiquitinates LC3B to increase LC3B levels and autophagic activity.
Jia R, Bonifacino JS
The Journal of biological chemistry 2021 Jan-Jun;296:100405
The Journal of biological chemistry 2021 Jan-Jun;296:100405
The Pah-R261Q mouse reveals oxidative stress associated with amyloid-like hepatic aggregation of mutant phenylalanine hydroxylase.
Aubi O, Prestegård KS, Jung-Kc K, Shi TS, Ying M, Grindheim AK, Scherer T, Ulvik A, McCann A, Spriet E, Thöny B, Martinez A
Nature communications 2021 Apr 6;12(1):2073
Nature communications 2021 Apr 6;12(1):2073
The degradation-promoting roles of deubiquitinases Ubp6 and Ubp3 in cytosolic and ER protein quality control.
Wu H, Ng DTW, Cheong I, Matsudaira P
PloS one 2020;15(5):e0232755
PloS one 2020;15(5):e0232755
Genetic Selection Based on a Ste6(*)C-HA-Ura3 Substrate Identifies New Cytosolic Quality Control Alleles in Saccharomyces cerevisiae.
Chan SN, Prasad R, Matsudaira P
G3 (Bethesda, Md.) 2020 Jun 1;10(6):1879-1891
G3 (Bethesda, Md.) 2020 Jun 1;10(6):1879-1891
IL-1β suppression of VE-cadherin transcription underlies sepsis-induced inflammatory lung injury.
Xiong S, Hong Z, Huang LS, Tsukasaki Y, Nepal S, Di A, Zhong M, Wu W, Ye Z, Gao X, Rao GN, Mehta D, Rehman J, Malik AB
The Journal of clinical investigation 2020 Jul 1;130(7):3684-3698
The Journal of clinical investigation 2020 Jul 1;130(7):3684-3698
Negative regulation of autophagy by UBA6-BIRC6-mediated ubiquitination of LC3.
Jia R, Bonifacino JS
eLife 2019 Nov 6;8
eLife 2019 Nov 6;8
Replication of multiple system atrophy prions in primary astrocyte cultures from transgenic mice expressing human α-synuclein.
Krejciova Z, Carlson GA, Giles K, Prusiner SB
Acta neuropathologica communications 2019 May 20;7(1):81
Acta neuropathologica communications 2019 May 20;7(1):81
APOPT1/COA8 assists COX assembly and is oppositely regulated by UPS and ROS.
Signes A, Cerutti R, Dickson AS, Benincá C, Hinchy EC, Ghezzi D, Carrozzo R, Bertini E, Murphy MP, Nathan JA, Viscomi C, Fernandez-Vizarra E, Zeviani M
EMBO molecular medicine 2019 Jan;11(1)
EMBO molecular medicine 2019 Jan;11(1)
A Mycobacterium tuberculosis surface protein recruits ubiquitin to trigger host xenophagy.
Chai Q, Wang X, Qiang L, Zhang Y, Ge P, Lu Z, Zhong Y, Li B, Wang J, Zhang L, Zhou D, Li W, Dong W, Pang Y, Gao GF, Liu CH
Nature communications 2019 Apr 29;10(1):1973
Nature communications 2019 Apr 29;10(1):1973
Ubiquitin-specific peptidase 2a (USP2a) deubiquitinates and stabilizes β-catenin.
Kim J, Alavi Naini F, Sun Y, Ma L
American journal of cancer research 2018;8(9):1823-1836
American journal of cancer research 2018;8(9):1823-1836
Cardiac-specific ablation of the E3 ubiquitin ligase Mdm2 leads to oxidative stress, broad mitochondrial deficiency and early death.
Hauck L, Stanley-Hasnain S, Fung A, Grothe D, Rao V, Mak TW, Billia F
PloS one 2017;12(12):e0189861
PloS one 2017;12(12):e0189861
Cycloheximide promotes paraptosis induced by inhibition of cyclophilins in glioblastoma multiforme.
Wang L, Gundelach JH, Bram RJ
Cell death & disease 2017 May 18;8(5):e2807
Cell death & disease 2017 May 18;8(5):e2807
Regulation of the ubiquitylation and deubiquitylation of CREB-binding protein modulates histone acetylation and lung inflammation.
Wei J, Dong S, Bowser RK, Khoo A, Zhang L, Jacko AM, Zhao Y, Zhao J
Science signaling 2017 Jun 13;10(483)
Science signaling 2017 Jun 13;10(483)
Small chaperons and autophagy protected neurons from necrotic cell death.
Lei Y, Liu K, Hou L, Ding L, Li Y, Liu L
Scientific reports 2017 Jul 18;7(1):5650
Scientific reports 2017 Jul 18;7(1):5650
Fbxl18 targets LRRK2 for proteasomal degradation and attenuates cell toxicity.
Ding X, Barodia SK, Ma L, Goldberg MS
Neurobiology of disease 2017 Feb;98:122-136
Neurobiology of disease 2017 Feb;98:122-136
Post-translational modifications of Annexin A2 are linked to its association with perinuclear nonpolysomal mRNP complexes.
Aukrust I, Rosenberg LA, Ankerud MM, Bertelsen V, Hollås H, Saraste J, Grindheim AK, Vedeler A
FEBS open bio 2017 Feb;7(2):160-173
FEBS open bio 2017 Feb;7(2):160-173
The E3 ligase Mule protects the heart against oxidative stress and mitochondrial dysfunction through Myc-dependent inactivation of Pgc-1α and Pink1.
Dadson K, Hauck L, Hao Z, Grothe D, Rao V, Mak TW, Billia F
Scientific reports 2017 Feb 2;7:41490
Scientific reports 2017 Feb 2;7:41490
Drosophila Mitf regulates the V-ATPase and the lysosomal-autophagic pathway.
Bouché V, Espinosa AP, Leone L, Sardiello M, Ballabio A, Botas J
Autophagy 2016;12(3):484-98
Autophagy 2016;12(3):484-98
Comparison of mHTT Antibodies in Huntington's Disease Mouse Models Reveal Specific Binding Profiles and Steady-State Ubiquitin Levels with Disease Development.
Bayram-Weston Z, Jones L, Dunnett SB, Brooks SP
PloS one 2016;11(5):e0155834
PloS one 2016;11(5):e0155834
Aberrant Splicing Promotes Proteasomal Degradation of L-type Ca(V)1.2 Calcium Channels by Competitive Binding for Ca(V)β Subunits in Cardiac Hypertrophy.
Hu Z, Wang JW, Yu D, Soon JL, de Kleijn DP, Foo R, Liao P, Colecraft HM, Soong TW
Scientific reports 2016 Oct 12;6:35247
Scientific reports 2016 Oct 12;6:35247
Multiple myeloma-derived MMP-13 mediates osteoclast fusogenesis and osteolytic disease.
Fu J, Li S, Feng R, Ma H, Sabeh F, Roodman GD, Wang J, Robinson S, Guo XE, Lund T, Normolle D, Mapara MY, Weiss SJ, Lentzsch S
The Journal of clinical investigation 2016 May 2;126(5):1759-72
The Journal of clinical investigation 2016 May 2;126(5):1759-72
Reactive oxygen species exert opposite effects on Tyr23 phosphorylation of the nuclear and cortical pools of annexin A2.
Grindheim AK, Hollås H, Raddum AM, Saraste J, Vedeler A
Journal of cell science 2016 Jan 15;129(2):314-28
Journal of cell science 2016 Jan 15;129(2):314-28
NQO1 inhibits proteasome-mediated degradation of HIF-1α.
Oh ET, Kim JW, Kim JM, Kim SJ, Lee JS, Hong SS, Goodwin J, Ruthenborg RJ, Jung MG, Lee HJ, Lee CH, Park ES, Kim C, Park HJ
Nature communications 2016 Dec 14;7:13593
Nature communications 2016 Dec 14;7:13593
Optineurin deficiency in mice is associated with increased sensitivity to Salmonella but does not affect proinflammatory NF-κB signaling.
Slowicka K, Vereecke L, Mc Guire C, Sze M, Maelfait J, Kolpe A, Saelens X, Beyaert R, van Loo G
European journal of immunology 2016 Apr;46(4):971-80
European journal of immunology 2016 Apr;46(4):971-80
Phosphorylation of OPTN by TBK1 enhances its binding to Ub chains and promotes selective autophagy of damaged mitochondria.
Richter B, Sliter DA, Herhaus L, Stolz A, Wang C, Beli P, Zaffagnini G, Wild P, Martens S, Wagner SA, Youle RJ, Dikic I
Proceedings of the National Academy of Sciences of the United States of America 2016 Apr 12;113(15):4039-44
Proceedings of the National Academy of Sciences of the United States of America 2016 Apr 12;113(15):4039-44
mRNA 3'-UTR shortening is a molecular signature of mTORC1 activation.
Chang JW, Zhang W, Yeh HS, de Jong EP, Jun S, Kim KH, Bae SS, Beckman K, Hwang TH, Kim KS, Kim DH, Griffin TJ, Kuang R, Yong J
Nature communications 2015 Jun 15;6:7218
Nature communications 2015 Jun 15;6:7218
The cell cycle regulator 14-3-3σ opposes and reverses cancer metabolic reprogramming.
Phan L, Chou PC, Velazquez-Torres G, Samudio I, Parreno K, Huang Y, Tseng C, Vu T, Gully C, Su CH, Wang E, Chen J, Choi HH, Fuentes-Mattei E, Shin JH, Shiang C, Grabiner B, Blonska M, Skerl S, Shao Y, Cody D, Delacerda J, Kingsley C, Webb D, Carlock C, Zhou Z, Hsieh YC, Lee J, Elliott A, Ramirez M, Bankson J, Hazle J, Wang Y, Li L, Weng S, Rizk N, Wen YY, Lin X, Wang H, Wang H, Zhang A, Xia X, Wu Y, Habra M, Yang W, Pusztai L, Yeung SC, Lee MH
Nature communications 2015 Jul 16;6:7530
Nature communications 2015 Jul 16;6:7530
Identification of a subnuclear body involved in sequence-specific cytokine RNA processing.
Lee S, Lee TA, Lee E, Kang S, Park A, Kim SW, Park HJ, Yoon JH, Ha SJ, Park T, Lee JS, Cheon JH, Park B
Nature communications 2015 Jan 5;6:5791
Nature communications 2015 Jan 5;6:5791
SLAP displays tumour suppressor functions in colorectal cancer via destabilization of the SRC substrate EPHA2.
Naudin C, Sirvent A, Leroy C, Larive R, Simon V, Pannequin J, Bourgaux JF, Pierre J, Robert B, Hollande F, Roche S
Nature communications 2014;5:3159
Nature communications 2014;5:3159
CSN6 drives carcinogenesis by positively regulating Myc stability.
Chen J, Shin JH, Zhao R, Phan L, Wang H, Xue Y, Post SM, Ho Choi H, Chen JS, Wang E, Zhou Z, Tseng C, Gully C, Velazquez-Torres G, Fuentes-Mattei E, Yeung G, Qiao Y, Chou PC, Su CH, Hsieh YC, Hsu SL, Ohshiro K, Shaikenov T, Wang H, Yeung SC, Lee MH
Nature communications 2014 Nov 14;5:5384
Nature communications 2014 Nov 14;5:5384
Ubiquitin-dependent regulation of MEKK2/3-MEK5-ERK5 signaling module by XIAP and cIAP1.
Takeda AN, Oberoi-Khanuja TK, Glatz G, Schulenburg K, Scholz RP, Carpy A, Macek B, Remenyi A, Rajalingam K
The EMBO journal 2014 Aug 18;33(16):1784-801
The EMBO journal 2014 Aug 18;33(16):1784-801
Monoclonal antibody against the ectodomain of E-cadherin (DECMA-1) suppresses breast carcinogenesis: involvement of the HER/PI3K/Akt/mTOR and IAP pathways.
Brouxhon SM, Kyrkanides S, Teng X, Raja V, O'Banion MK, Clarke R, Byers S, Silberfeld A, Tornos C, Ma L
Clinical cancer research : an official journal of the American Association for Cancer Research 2013 Jun 15;19(12):3234-46
Clinical cancer research : an official journal of the American Association for Cancer Research 2013 Jun 15;19(12):3234-46
Drosophila Fip200 is an essential regulator of autophagy that attenuates both growth and aging.
Kim M, Park HL, Park HW, Ro SH, Nam SG, Reed JM, Guan JL, Lee JH
Autophagy 2013 Aug;9(8):1201-13
Autophagy 2013 Aug;9(8):1201-13
Dissecting the pathways that destabilize mutant p53: the proteasome or autophagy?
Choudhury S, Kolukula VK, Preet A, Albanese C, Avantaggiati ML
Cell cycle (Georgetown, Tex.) 2013 Apr 1;12(7):1022-9
Cell cycle (Georgetown, Tex.) 2013 Apr 1;12(7):1022-9
An intrinsically disordered region of the acetyltransferase p300 with similarity to prion-like domains plays a role in aggregation.
Kirilyuk A, Shimoji M, Catania J, Sahu G, Pattabiraman N, Giordano A, Albanese C, Mocchetti I, Toretsky JA, Uversky VN, Avantaggiati ML
PloS one 2012;7(11):e48243
PloS one 2012;7(11):e48243
Mitochondrial division ensures the survival of postmitotic neurons by suppressing oxidative damage.
Kageyama Y, Zhang Z, Roda R, Fukaya M, Wakabayashi J, Wakabayashi N, Kensler TW, Reddy PH, Iijima M, Sesaki H
The Journal of cell biology 2012 May 14;197(4):535-51
The Journal of cell biology 2012 May 14;197(4):535-51
Analysis of nuclear factor-κB (NF-κB) essential modulator (NEMO) binding to linear and lysine-linked ubiquitin chains and its role in the activation of NF-κB.
Kensche T, Tokunaga F, Ikeda F, Goto E, Iwai K, Dikic I
The Journal of biological chemistry 2012 Jul 6;287(28):23626-34
The Journal of biological chemistry 2012 Jul 6;287(28):23626-34
Gene expression levels assessed by CA1 pyramidal neuron and regional hippocampal dissections in Alzheimer's disease.
Ginsberg SD, Alldred MJ, Che S
Neurobiology of disease 2012 Jan;45(1):99-107
Neurobiology of disease 2012 Jan;45(1):99-107
Optineurin is colocalized with ubiquitin in Marinesco bodies.
Schwab C, Yu S, McGeer PL
Acta neuropathologica 2012 Feb;123(2):289-92
Acta neuropathologica 2012 Feb;123(2):289-92
Dietary downregulation of mutant p53 levels via glucose restriction: mechanisms and implications for tumor therapy.
Rodriguez OC, Choudhury S, Kolukula V, Vietsch EE, Catania J, Preet A, Reynoso K, Bargonetti J, Wellstein A, Albanese C, Avantaggiati ML
Cell cycle (Georgetown, Tex.) 2012 Dec 1;11(23):4436-46
Cell cycle (Georgetown, Tex.) 2012 Dec 1;11(23):4436-46
Subunit 6 of the COP9 signalosome promotes tumorigenesis in mice through stabilization of MDM2 and is upregulated in human cancers.
Zhao R, Yeung SC, Chen J, Iwakuma T, Su CH, Chen B, Qu C, Zhang F, Chen YT, Lin YL, Lee DF, Jin F, Zhu R, Shaikenov T, Sarbassov D, Sahin A, Wang H, Wang H, Lai CC, Tsai FJ, Lozano G, Lee MH
The Journal of clinical investigation 2011 Mar;121(3):851-65
The Journal of clinical investigation 2011 Mar;121(3):851-65
Synaptic dysbindin-1 reductions in schizophrenia occur in an isoform-specific manner indicating their subsynaptic location.
Talbot K, Louneva N, Cohen JW, Kazi H, Blake DJ, Arnold SE
PloS one 2011 Mar 1;6(3):e16886
PloS one 2011 Mar 1;6(3):e16886
Protein interactions among Fe65, the low-density lipoprotein receptor-related protein, and the amyloid precursor protein.
Mulvihill MM, Guttman M, Komives EA
Biochemistry 2011 Jul 19;50(28):6208-16
Biochemistry 2011 Jul 19;50(28):6208-16
A photoconvertible fluorescent reporter to track chaperone-mediated autophagy.
Koga H, Martinez-Vicente M, Macian F, Verkhusha VV, Cuervo AM
Nature communications 2011 Jul 12;2:386
Nature communications 2011 Jul 12;2:386
Amyloid precursor-like protein 2 cleavage contributes to neuronal intranuclear inclusions and cytotoxicity in spinocerebellar ataxia-7 (SCA7).
Takahashi-Fujigasaki J, Breidert T, Fujigasaki H, Duyckaerts C, Camonis JH, Brice A, Lebre AS
Neurobiology of disease 2011 Jan;41(1):33-42
Neurobiology of disease 2011 Jan;41(1):33-42
SCA1-like disease in mice expressing wild-type ataxin-1 with a serine to aspartic acid replacement at residue 776.
Duvick L, Barnes J, Ebner B, Agrawal S, Andresen M, Lim J, Giesler GJ, Zoghbi HY, Orr HT
Neuron 2010 Sep 23;67(6):929-35
Neuron 2010 Sep 23;67(6):929-35
Neurodegeneration by polyglutamine Atrophin is not rescued by induction of autophagy.
Nisoli I, Chauvin JP, Napoletano F, Calamita P, Zanin V, Fanto M, Charroux B
Cell death and differentiation 2010 Oct;17(10):1577-87
Cell death and differentiation 2010 Oct;17(10):1577-87
Psh1 is an E3 ubiquitin ligase that targets the centromeric histone variant Cse4.
Hewawasam G, Shivaraju M, Mattingly M, Venkatesh S, Martin-Brown S, Florens L, Workman JL, Gerton JL
Molecular cell 2010 Nov 12;40(3):444-54
Molecular cell 2010 Nov 12;40(3):444-54
Different modes of ubiquitination of the adaptor TRAF3 selectively activate the expression of type I interferons and proinflammatory cytokines.
Tseng PH, Matsuzawa A, Zhang W, Mino T, Vignali DA, Karin M
Nature immunology 2010 Jan;11(1):70-5
Nature immunology 2010 Jan;11(1):70-5
Chemical genetics approach to restoring p27Kip1 reveals novel compounds with antiproliferative activity in prostate cancer cells.
Rico-Bautista E, Yang CC, Lu L, Roth GP, Wolf DA
BMC biology 2010 Dec 23;8:153
BMC biology 2010 Dec 23;8:153
Autophagy in load-induced heart disease.
Zhu H, Rothermel BA, Hill JA
Methods in enzymology 2009;453:343-63
Methods in enzymology 2009;453:343-63
Interactions of the NPXY microdomains of the low density lipoprotein receptor-related protein 1.
Guttman M, Betts GN, Barnes H, Ghassemian M, van der Geer P, Komives EA
Proteomics 2009 Nov;9(22):5016-28
Proteomics 2009 Nov;9(22):5016-28
Early-onset and robust amyloid pathology in a new homozygous mouse model of Alzheimer's disease.
Willuweit A, Velden J, Godemann R, Manook A, Jetzek F, Tintrup H, Kauselmann G, Zevnik B, Henriksen G, Drzezga A, Pohlner J, Schoor M, Kemp JA, von der Kammer H
PloS one 2009 Nov 20;4(11):e7931
PloS one 2009 Nov 20;4(11):e7931
Thioredoxin-related Protein 32 is an arsenite-regulated Thiol Reductase of the proteasome 19 S particle.
Wiseman RL, Chin KT, Haynes CM, Stanhill A, Xu CF, Roguev A, Krogan NJ, Neubert TA, Ron D
The Journal of biological chemistry 2009 May 29;284(22):15233-45
The Journal of biological chemistry 2009 May 29;284(22):15233-45
IKK phosphorylates Huntingtin and targets it for degradation by the proteasome and lysosome.
Thompson LM, Aiken CT, Kaltenbach LS, Agrawal N, Illes K, Khoshnan A, Martinez-Vincente M, Arrasate M, O'Rourke JG, Khashwji H, Lukacsovich T, Zhu YZ, Lau AL, Massey A, Hayden MR, Zeitlin SO, Finkbeiner S, Green KN, LaFerla FM, Bates G, Huang L, Patterson PH, Lo DC, Cuervo AM, Marsh JL, Steffan JS
The Journal of cell biology 2009 Dec 28;187(7):1083-99
The Journal of cell biology 2009 Dec 28;187(7):1083-99
Regulation of mitochondrial morphology by USP30, a deubiquitinating enzyme present in the mitochondrial outer membrane.
Nakamura N, Hirose S
Molecular biology of the cell 2008 May;19(5):1903-11
Molecular biology of the cell 2008 May;19(5):1903-11
The NER protein Rad33 shows functional homology to human Centrin2 and is involved in modification of Rad4.
den Dulk B, van Eijk P, de Ruijter M, Brandsma JA, Brouwer J
DNA repair 2008 Jun 1;7(6):858-68
DNA repair 2008 Jun 1;7(6):858-68
Rapid identification of differentiation markers from whole epithelial cells by matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry and statistical analysis.
Marvin-Guy LF, Duncan P, Wagnière S, Antille N, Porta N, Affolter M, Kussmann M
Rapid communications in mass spectrometry : RCM 2008 Apr;22(8):1099-108
Rapid communications in mass spectrometry : RCM 2008 Apr;22(8):1099-108
Geldanamycin induces G2 arrest in U87MG glioblastoma cells through downregulation of Cdc2 and cyclin B1.
Nomura N, Nomura M, Newcomb EW, Zagzag D
Biochemical pharmacology 2007 May 15;73(10):1528-36
Biochemical pharmacology 2007 May 15;73(10):1528-36
PDLIM2-mediated termination of transcription factor NF-kappaB activation by intranuclear sequestration and degradation of the p65 subunit.
Tanaka T, Grusby MJ, Kaisho T
Nature immunology 2007 Jun;8(6):584-91
Nature immunology 2007 Jun;8(6):584-91
Kinase MEKK1 is required for CD40-dependent activation of the kinases Jnk and p38, germinal center formation, B cell proliferation and antibody production.
Gallagher E, Enzler T, Matsuzawa A, Anzelon-Mills A, Otero D, Holzer R, Janssen E, Gao M, Karin M
Nature immunology 2007 Jan;8(1):57-63
Nature immunology 2007 Jan;8(1):57-63
Allosteric activation of human glucokinase by free polyubiquitin chains and its ubiquitin-dependent cotranslational proteasomal degradation.
Bjørkhaug L, Molnes J, Søvik O, Njølstad PR, Flatmark T
The Journal of biological chemistry 2007 Aug 3;282(31):22757-64
The Journal of biological chemistry 2007 Aug 3;282(31):22757-64
RhoB facilitates c-Myc turnover by supporting efficient nuclear accumulation of GSK-3.
Huang M, Kamasani U, Prendergast GC
Oncogene 2006 Mar 2;25(9):1281-9
Oncogene 2006 Mar 2;25(9):1281-9
Activation of the E3 ubiquitin ligase Itch through a phosphorylation-induced conformational change.
Gallagher E, Gao M, Liu YC, Karin M
Proceedings of the National Academy of Sciences of the United States of America 2006 Feb 7;103(6):1717-22
Proceedings of the National Academy of Sciences of the United States of America 2006 Feb 7;103(6):1717-22
Experimental tests to definitively determine ubiquitylation of a substrate.
Bloom J, Pagano M
Methods in enzymology 2005;399:249-66
Methods in enzymology 2005;399:249-66
Defining the role of ubiquitin-interacting motifs in the polyglutamine disease protein, ataxin-3.
Berke SJ, Chai Y, Marrs GL, Wen H, Paulson HL
The Journal of biological chemistry 2005 Sep 9;280(36):32026-34
The Journal of biological chemistry 2005 Sep 9;280(36):32026-34
Geldanamycin-induced degradation of Chk1 is mediated by proteasome.
Nomura M, Nomura N, Yamashita J
Biochemical and biophysical research communications 2005 Sep 30;335(3):900-5
Biochemical and biophysical research communications 2005 Sep 30;335(3):900-5
Methylation silencing of SOCS-3 promotes cell growth and migration by enhancing JAK/STAT and FAK signalings in human hepatocellular carcinoma.
Niwa Y, Kanda H, Shikauchi Y, Saiura A, Matsubara K, Kitagawa T, Yamamoto J, Kubo T, Yoshikawa H
Oncogene 2005 Sep 22;24(42):6406-17
Oncogene 2005 Sep 22;24(42):6406-17
Involvement of the p97-Ufd1-Npl4 complex in the regulated endoplasmic reticulum-associated degradation of inositol 1,4,5-trisphosphate receptors.
Alzayady KJ, Panning MM, Kelley GG, Wojcikiewicz RJ
The Journal of biological chemistry 2005 Oct 14;280(41):34530-7
The Journal of biological chemistry 2005 Oct 14;280(41):34530-7
Role of p97 AAA-ATPase in the retrotranslocation of the cholera toxin A1 chain, a non-ubiquitinated substrate.
Kothe M, Ye Y, Wagner JS, De Luca HE, Kern E, Rapoport TA, Lencer WI
The Journal of biological chemistry 2005 Jul 29;280(30):28127-32
The Journal of biological chemistry 2005 Jul 29;280(30):28127-32
Histomathematical analysis of clinical specimens: challenges and progress.
Gannot G, Gillespie JW, Chuaqui RF, Tangrea MA, Linehan WM, Emmert-Buck MR
The journal of histochemistry and cytochemistry : official journal of the Histochemistry Society 2005 Feb;53(2):177-85
The journal of histochemistry and cytochemistry : official journal of the Histochemistry Society 2005 Feb;53(2):177-85
Similar patterns of mitochondrial vulnerability and rescue induced by genetic modification of alpha-synuclein, parkin, and DJ-1 in Caenorhabditis elegans.
Ved R, Saha S, Westlund B, Perier C, Burnam L, Sluder A, Hoener M, Rodrigues CM, Alfonso A, Steer C, Liu L, Przedborski S, Wolozin B
The Journal of biological chemistry 2005 Dec 30;280(52):42655-42668
The Journal of biological chemistry 2005 Dec 30;280(52):42655-42668
Proteasome mediates dopaminergic neuronal degeneration, and its inhibition causes alpha-synuclein inclusions.
Sawada H, Kohno R, Kihara T, Izumi Y, Sakka N, Ibi M, Nakanishi M, Nakamizo T, Yamakawa K, Shibasaki H, Yamamoto N, Akaike A, Inden M, Kitamura Y, Taniguchi T, Shimohama S
The Journal of biological chemistry 2004 Mar 12;279(11):10710-9
The Journal of biological chemistry 2004 Mar 12;279(11):10710-9
Distinct steps in dislocation of luminal endoplasmic reticulum-associated degradation substrates: roles of endoplamic reticulum-bound p97/Cdc48p and proteasome.
Elkabetz Y, Shapira I, Rabinovich E, Bar-Nun S
The Journal of biological chemistry 2004 Feb 6;279(6):3980-9
The Journal of biological chemistry 2004 Feb 6;279(6):3980-9
A signalling pathway controlling c-Myc degradation that impacts oncogenic transformation of human cells.
Yeh E, Cunningham M, Arnold H, Chasse D, Monteith T, Ivaldi G, Hahn WC, Stukenberg PT, Shenolikar S, Uchida T, Counter CM, Nevins JR, Means AR, Sears R
Nature cell biology 2004 Apr;6(4):308-18
Nature cell biology 2004 Apr;6(4):308-18
Conformational change and mitochondrial translocation of Bax accompany proteasome inhibitor-induced apoptosis of chronic lymphocytic leukemic cells.
Dewson G, Snowden RT, Almond JB, Dyer MJ, Cohen GM
Oncogene 2003 May 1;22(17):2643-54
Oncogene 2003 May 1;22(17):2643-54
Acquired FANCA dysfunction and cytogenetic instability in adult acute myelogenous leukemia.
Lensch MW, Tischkowitz M, Christianson TA, Reifsteck CA, Speckhart SA, Jakobs PM, O'Dwyer ME, Olson SB, Le Beau MM, Hodgson SV, Mathew CG, Larson RA, Bagby GC Jr
Blood 2003 Jul 1;102(1):7-16
Blood 2003 Jul 1;102(1):7-16
blue cheese mutations define a novel, conserved gene involved in progressive neural degeneration.
Finley KD, Edeen PT, Cumming RC, Mardahl-Dumesnil MD, Taylor BJ, Rodriguez MH, Hwang CE, Benedetti M, McKeown M
The Journal of neuroscience : the official journal of the Society for Neuroscience 2003 Feb 15;23(4):1254-64
The Journal of neuroscience : the official journal of the Society for Neuroscience 2003 Feb 15;23(4):1254-64
Ubiquitylation of BAG-1 suggests a novel regulatory mechanism during the sorting of chaperone substrates to the proteasome.
Alberti S, Demand J, Esser C, Emmerich N, Schild H, Hohfeld J
The Journal of biological chemistry 2002 Nov 29;277(48):45920-7
The Journal of biological chemistry 2002 Nov 29;277(48):45920-7
Immunohistochemical study of p53-associated proteins in rat brain following lithium-pilocarpine status epilepticus.
Tan Z, Sankar R, Tu W, Shin D, Liu H, Wasterlain CG, Schreiber SS
Brain research 2002 Mar 1;929(1):129-38
Brain research 2002 Mar 1;929(1):129-38
Inhibition of Src family kinases blocks epidermal growth factor (EGF)-induced activation of Akt, phosphorylation of c-Cbl, and ubiquitination of the EGF receptor.
Kassenbrock CK, Hunter S, Garl P, Johnson GL, Anderson SM
The Journal of biological chemistry 2002 Jul 12;277(28):24967-75
The Journal of biological chemistry 2002 Jul 12;277(28):24967-75
Interferon-gamma-mediated activation and ubiquitin-proteasome-dependent degradation of PPARgamma in adipocytes.
Floyd ZE, Stephens JM
The Journal of biological chemistry 2002 Feb 8;277(6):4062-8
The Journal of biological chemistry 2002 Feb 8;277(6):4062-8
Regulation of the ubiquitin-conjugating enzyme hHR6A by CDK-mediated phosphorylation.
Sarcevic B, Mawson A, Baker RT, Sutherland RL
The EMBO journal 2002 Apr 15;21(8):2009-18
The EMBO journal 2002 Apr 15;21(8):2009-18
Cancer-predisposing mutations within the RING domain of BRCA1: loss of ubiquitin protein ligase activity and protection from radiation hypersensitivity.
Ruffner H, Joazeiro CA, Hemmati D, Hunter T, Verma IM
Proceedings of the National Academy of Sciences of the United States of America 2001 Apr 24;98(9):5134-9
Proceedings of the National Academy of Sciences of the United States of America 2001 Apr 24;98(9):5134-9
Proteasomes regulate erythropoietin receptor and signal transducer and activator of transcription 5 (STAT5) activation. Possible involvement of the ubiquitinated Cis protein.
Verdier F, Chrétien S, Muller O, Varlet P, Yoshimura A, Gisselbrecht S, Lacombe C, Mayeux P
The Journal of biological chemistry 1998 Oct 23;273(43):28185-90
The Journal of biological chemistry 1998 Oct 23;273(43):28185-90
The Ras-specific exchange factors mouse Sos1 (mSos1) and mSos2 are regulated differently: mSos2 contains ubiquitination signals absent in mSos1.
Nielsen KH, Papageorge AG, Vass WC, Willumsen BM, Lowy DR
Molecular and cellular biology 1997 Dec;17(12):7132-8
Molecular and cellular biology 1997 Dec;17(12):7132-8
Identification of a neural alpha-catenin as a key regulator of cadherin function and multicellular organization.
Hirano S, Kimoto N, Shimoyama Y, Hirohashi S, Takeichi M
Cell 1992 Jul 24;70(2):293-301
Cell 1992 Jul 24;70(2):293-301
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Western blot analysis of ubiquitin was performed by loading 30 µg of cell lysate of mouse thymocytes treated with 0 µM MG132 for 5 hr at 37C (lane 1) and cell lysate of mouse thymocytes treated with 20 µM MG132 for 5 hr at 37C (lane 2) in 1xSDS sample buffer and SeeBlue® Plus2 Pre-stained Protein Standard (Product # LC5925) onto a NuPAGE™ Novex™ 4-12% Bis-Tris Protein Gels (Product # NP0324BOX). Proteins were transferred to nitrocellulose membrane. Membrane was blocked in 1xDPBS (Product # 14190-136) containing 0.1% Tween 20 and 5% non-fat milk for one hour at room temperature. Ubiquitin was detected at numerous molecular weights using a Ubiquitin antibody (Product # 13-1600) at a dilution of 1:1,000 in 1xDPBS (Product # 14190-136) containing 0.1% Tween 20 and 5% non-fat milk overnight at 4°C on a rocking platform, followed by a Goat anti-Mouse IgG-HRP secondary antibody at a dilution of 1:5,000 for one hour at room temperature. Chemiluminescent detection was performed using Immoblion Western and the ImageQuant LAS4000 imager. Data Courtesy of Dr. Zhi Wen at the Univeristy of Wisconsin, Madison.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescent staining of the superior frontal cortex of Huntington's disease tissue with Ms anti-Ubiquitin (Product # 13-1600). Image reproduced from Holbert S, et al. PNAS 2003.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescent staining of the superior frontal cortex of Huntington's disease tissue with Ms anti-Ubiquitin (Product # 13-1600). Image reproduced from Holbert S, et al. PNAS 2003.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
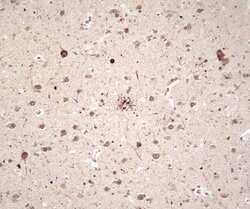
- Experimental details
- Immunohistochemical staining of Alzheimer's disease brain tissue using Ms anti-Ubiquitin (Product # 13-1600).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
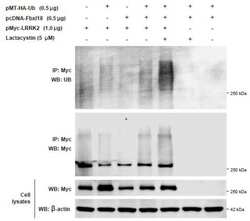
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5 Validation of SMIP001 and SMIP004 . (a) Chemical structure of the two small molecule inhibitors of p27 depletions (SMIPs) identified in the screen. (b) Cell lysates from LNCaP-S14 treated with 40 muM SMIPs for 24 h were analysed by immunoblotting. Roscovitine (R) was used as positive control. The data shown is representative of two independent experiments. (c) LNCaP-S14 cells were treated with increasing concentrations of SMIPs (5-40 muM) for 18 h followed by fixation and staining for p27. The graph represents data from 24 replicate wells per concentration. (d) LNCaP-S14 cells were treated with SMIPs (5-40 muM) for 18 h and analysed by immunoblotting for the levels of p27 and p21. (e) p27, p21 and SKP2 stability in LNCaP-S14 cells treated with 40 muM SMIPs for 18 h. Cycloheximide (CHX, 100 mug/mL) was added for the indicated time points prior to preparation of cell lysates and immunoblot analysis. MG132 was used as positive control. The graphs represent the quantification of p27, p21 and SKP2 levels relative to the zero hour time point. The data is representative of three independent experiments. (f) LNCaP-S14 cells were treated with SMIP00s (40 muM) for 18 h followed by extraction of total RNA. The mRNA levels of p27, p21 and SKP2 were quantified by quantitative polymerase chain reaction as described in the Methods section. All data was normalized to GAPDH and is expressed as fold induction. The graph is representative of two independent experiments. (g) Total lysat
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 1 p300 localizes in aggresomes. A-C . Cos7 cells were treated with DMSO (A) or treated with 5 uM MG132 for 16 hours (B-C) and cells were stained with a monoclonal mix of anti-ubiquitin (A and B), the anti-p300 polyclonal C20 antibody (A-C), or anti-vimentin monoclonal (C) antibodies (see materials and methods for details on the antibodies). The arrow in panel A, indicates the presence of p300 in diffused cytoplasmic aggregates. The arrows in B and C indicate the position of representative aggresomes, enclosed by vimentin and containing ubiquitin. Cells were subjected to fluorescence deconvolution and image reconstruction by using a Zeiss microscope (Axiovert 200M) with deconvolution capabilities (Axiovision 4.1). The merge panels represent the deconvoluted images from the Alexa-Fluor 488 (green), Alexa-Fluor 568 (red), and DAPI signals. D. Quantification of the immuno-fluorescence experiments. The percentage of cells containing aggresomes (black bars) and of aggresomes containing p300 (gray bars) was calculated in mock treated (-) or MG132 treated (+) cells. E. Cos7 cells were mock treated (-; lanes 1 and 4) or treated with MG132 (+; lanes 2, 3 and 5) and cell extracts were subjected to immuno-precipitation with a control antibody (lane 3) or with the anti-p300 antibody (lanes 4 and 5). Immuno-precipitation reactions were divided into different aliquots and subjected to immuno-blot with anti p300 (top panel), anti-HDAC6 (mid panel) or anti dynein antibodies (bottom pan
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
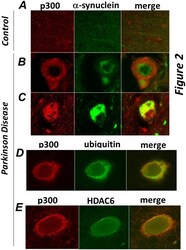
- Figure 2 p300 co-localizes with a-synuclein, ubiquitin and HDAC6 in brain of patients affected by Parkinson Disease. Sections of midbrains or cortex from normal control patients ( A ), or from patients by Parkinson Disease ( B-E ) were immuno-stained with the anti-p300 (red) and alpha-synuclein (green, panels B,C ), or with anti-ubiquitin (green, panel D), or with anti-HDAC6 (green, panel E) antibodies as indicated at the top of each panel. Different sections from the same patients or from different patients were subjected to analysis.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 6 The p300 PSPD is sufficient for aggresome localization. A-D. Cell transfected with the vector expressing RFP alone and treated with MG132 (panel A), or transfected with the RFP-PSPD (panels B-D) were mock treated (B), or treated with MG132 ( C-D ) and subsequently stained for vimentin (green). Panels C and D contain two representative fields of cells transfected with RFP-PSPD. Arrows indicate the position of aggregates formed in the absence (panel B) or in the presence of MG132 (C and D). E. Interaction of p300 with ubiquitinated protein species. The epitope-flagged vector expressing the PSPD (lanes 1-5), or the TAZ2 (lanes 6,7 and 10,11) or the ZZ (lanes 8,9 and 12,13) were transfected in Cos7 cells. Cells were left untreated (-) or treated with MG132 (+), cell extracts were immuno-precipitated with the anti-Flag antibody (lanes 1,2; and 6-to-9), or with a control isotype matched antibody (lane 3) and the products of these immuno-precipitation reactions were probed with ubiquitin antibodies as indicated at the bottom of each panel. Alternatively, the anti-Flag immuno-blot on total cell extracts shows the total amount of p300 proteins present in these reactions, which were derived from approximately 1/50 of total extracts.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 8 Effects of p300 on the misfolded protein response. A. A549 cells were transfected with scramble siRNA or with the p300 specific siRNA as described previously, and stained with the anti-ubiquitin (green) or anti-vimentin (red) antibody. The percentage of cells containing ubiquitin aggregates is quantified in panel B . C . A549 cells were transfected with the scramble- or p300-specific siRNAs were mock treated or treated with MG132 for 16 hours, then harvested for flow cytometry. Panel C shows the Propidium Iodide profiles, and percentages of cells in the G1 or G2 phases of the cell cycle are indicated at the top of each relative peak. D. A549 cells transfected with scramble (black bars) or with the p300 siRNAs (gray bars), were treated with MG132. In one set of samples MG132 was washed out after 12-14 hours of treatment (indicated as MG132 w/o) and cells were allowed to recover for about four days, at which time they were counted. Alternatively, cell growth was monitored for the same period of time in the presence of MG132 (indicated as MG132 on). Error bars represent standard deviations. E-F. Control (wt) or p300 mouse embryo fibroblasts (MEF) were mock treated or treated with MG132 for 16-24 hours (panel E), or alternatively, subjected to Heat Shock (HS, panel F) by incubating the cells at 40degC for two hours. Cells were allowed to recover from MG132 treatment or HS for 24-48 hours and were counted. Cell viability was assessed with trypan blue exclusion.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
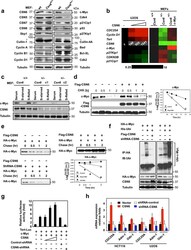
- Figure 1 Loss of CSN6 leads to destabilization of Myc and decreases Myc transcriptional activity (a) Immunoblot analysis of Csn6 +/- MEFs. Primary MEF cells were prepared from 13.5-day embryos derived from wild-type (wt) Csn6 +/+ and Csn6 +/- mice. After 4 days of culture, cell lysates were analyzed by indicated antibodies. (b) Quantitative RT-PCR analysis of mRNA levels for indicated c-Myc target genes in CSN6-transfected U2OS cells or Csn6 +/- MEFs. The quantitated mRNA expression level was normalized to GAPDH mRNA. Two MEFs were examined for each genotype. Expression level of transfected CSN6 is indicated as numbers on the heat map, and the heat map depicts the natural logarithm of fold-change in mRNA expression. (c) CSN6 knockdown abrogated serum-induced elevation of Myc. Csn6 +/+ MEFs and Csn6 +/- clone 1 (C1) and clone 2 (C2) MEFs were serum-starved for 24 hours and then were refed with serum for 24 hours. Lysates were analyzed by indicated antibodies. (d) CSN6 decreased Myc turnover. 293T cells were transfected with Flag-CSN6 and were then treated with cycloheximide (CHX; 100 ug/ml) for the indicated times. The immunoblot of Myc signal at each time point was measured using a densitometer and the integrated optical density of Myc was measured. The turnover of Myc is indicated graphically. (e) CSN6 expression affected Myc turnover. 35 S-pulse-labeled HA-Myc protein was immunoprecipitated from indicated transfected 293T cell lysates. The mixture was separated by sodium do
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5 14-3-3sigma inhibits cancer metabolism by promoting Myc degradation (a) Loss of 14-3-3sigma upregulated Myc-induced glycolytic genes. The expression of glycolytic genes in HCT116 WT (14-3-3sigma +/+ ) and HCT116 14-3-3sigma -/- cells were measured using Realtime PCR (left panel). Average+-95%CI, n=4, ANOVA,* P< 0.05. Western Blot was performed using the indicated antibodies (right panel). (b) 14-3-3sigma downregulated Myc-induced glycolytic genes. The mRNA expression of glycolytic genes were compared between indicated cells using Realtime PCR (left panel). Average+-95%CI, n=3, ANOVA,* P< 0.05. Western Blot was performed using the indicated antibodies (right panel). (c) 14-3-3sigma reduced expression of Myc-induced glutaminolytic genes. Realtime PCR and Western Blot were used to compare the expression of Myc-induced glutaminolytic genes at mRNA and protein levels. Average+-95%CI, n=3, ANOVA,* P< 0.05. (d) 14-3-3sigma suppressed TFAM expression and reduced MT-CO1/PPRC1 ratio in vitro. TFAM is an important Myc-induced mitochondrial transcription factor. MT-CO1 is a mitochondrial gene encoding cytochrome c oxidase. PPRC1 , peroxisome proliferator-activated receptor gamma, coactivator-related 1, is a gene of nuclear DNA. Average+-95% CI, n=3, Student t-test, * p < 0.05. (e) 14-3-3sigma increased Myc ubiquitination. Indicated cells were treated with Doxycycline to induce Flag-14-3-3sigma expression (+). Non-induced cells were used as a control (-). Cell lysates were immuno
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

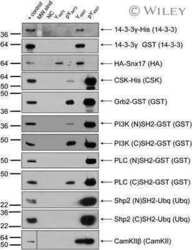
- Experimental details
- Fig. 5. Incorporation of ubiquitylated pTyr23AnxA2 into extracellular vesicles (exosomes) and decrease in the cortical pool of pTyr23AnxA2 after prolonged treatment of cells with H 2 O 2 . (A) PC12 cells were grown in exosome-depleted medium in the presence of H 2 O 2 for 0 min (lanes 1,5), 15 min (lanes 2,6), 1 h (lanes 3,7), or 2 h (lanes 4,8). ECM-bound proteins were released by EGTA (lanes 1-4), whereas extracellular vesicles were isolated from the culture medium by the ExoQuick-TC method (lanes 5-8). 100 ug of protein from the EGTA-released fractions (lanes 1-4) and the control extracellular vesicle fraction (lane 5), or an equal volume of extracellular vesicles from H 2 O 2 -treated cells (lanes 6-8) were separated by 10% SDS-PAGE, transferred to nitro-cellulose membranes and probed with antibodies against pTyr23AnxA2, total AnxA2, CD63, TSG-101 and T-cadherin, as indicated. (B) Following 1 h treatment of PC12 cells with 1 mM H 2 O 2 , proteins (600 ug) present in purified extracellular vesicles were immunoprecipitated (IP) by monoclonal AnxA2 antibodies (lane 1) after pre-clearance of the samples with normal mouse IgG (lane 2). The proteins were subjected to 10% SDS-PAGE and immunoblot analysis using monoclonal antibodies against pTyr23AnxA2 or ubiquitin by loading half of the immunoprecipitation sample on each gel. The bands representing ubiquitylated AnxA2 (square bracket; Ub-AnxA2) and IgG light chain (L C , arrowhead) are indicated to the right. (A,B) Following inc
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 1 Acute genetic deletion of Mule induces concentric hypertrophy with cardiac dysfunction and premature death. ( A ) Down-regulation of Mule mRNA levels in human end-stage heart failure determined by RT- qPCR. ( B ) To specifically inactivate Mule and Myc in the adult heart, we employed a tamoxifen (Tam) inducible Cre- loxP system in which Cre recombinase expression is controlled by the cardiomyocyte-specific myosin heavy chain 6 promoter ( mcm ). 4-hydroxytamoxifen (Tam) was injected intraperitoneally daily for four consecutive days in 10 week-old mice. The day of the last Tam injection was set as day zero. Veh, vehicle. Nt, no treatment. Immunoblot analysis of cytoplasmic Mule and nuclear Myc levels in left ventricular extracts (60 mug total protein/lane) of Mule fl / fl ( y ) ;mcm mice at 7 days post-Tam employing specific antibodies as indicated on the left. Animals were 12 weeks old at the time of analysis. For normalization, Western blots were probed with anti-tubulin (Tub) for cytoplasmic fractions, and anti-nucleophosmin (Npm1) for nuclear fractions. Immunoblots were repeated at least once with similar results. Transcript levels of Mule and Myc in Mule fl / fl ( y ) ;mcm mice at 7 d post-Tam as analyzed by RT-qPCR. n = 4. ( C ) Mule is indispensable for Myc protein stability in the heart by regulation of its ubiquitin-mediated proteasomal degradation. At 7 d post-Tam. Mule fl / fl ( y ) ;mcm , mice were intraperitoneally injected with the proteasomal inhibitor M
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 2 Pup2 predominantly affects CytoQC compared to ERAD. (A) Rescue of the CytoQC-defective phenotype observed with the exogenous expression of wild-type Pup2 ( PUP2 ) in the pup2 -10 mutant. Equal numbers of each strain were spotted as described in Figure 1C with SC-Trp-His and SC-Trp-Ura plates for selection. Vector: empty vector. (B) Missense mutation in spontaneous mutant pup2 -10 is present at residue 101, replacing Leucine for Proline. (C-D) Degradation kinetics of CytoQC and ERAD substrates were determined by pulse chase analyses. Strains were pulsed with 35S-Met/Cys for 5 min for Ste6 *C and Ste6 * and 10 min for ssPrA, CPY* and Sec61 -2, followed by chase for the indicated time points. (E) CytoQC substrate ssPrA is localized predominantly in the nucleus in WT and pup2 -10 . Substrates were detected with anti-HA antibody (green). The ER and nuclear envelope were visualized with anti- Kar2 antiserum (red). Nucleus was visualized with DAPI staining. Scale bar: 2 µm. (F) Accumulation of polyubiquitinated Ste6 *C and DeltassPrA was observed in pup2 -10 mutant compared to WT. Misfolded cytosolic substrates expressed in WT and pup2 -10 were immunoprecipitated (IP) by anti-HA antibody, resolved by SDS-PAGE and analyzed by immunoblotting (IB) with anti-ubiquitin antibody to detect polyubiquitinated substrates.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 5 Reduced free ubiquitin levels cannot fully account for degradation delays in doa4 . (A) Immunoblot of free ubiquitin levels in WT and doa4 deletion strains without any substrate, expressing CytoQC substrates Ste6 *C or ssPrA, and expressing either substrate in the presence of ubiquitin overexpression (Ub). Free ubiquitin was probed with the mouse anti-ubiquitin antibody and PGK was probed as loading control. Unique ubiquitin conjugates characteristic of doa4 are indicated by the black bracket. Graph shows average relative levels of free ubiquitin in doa4 compared to the corresponding WT. Error bars indicate standard deviation of the mean of three independent experiments. (B) Partial restoration of polyubiquitinated substrate levels in doa4 with the overexpression of ubiquitin (Ub). Misfolded CytoQC substrates expressed in WT and doa4 were processed and analyzed as described in Figure 2F . (C) Degradation delay of Ste6 *C and ssPrA was only partially suppressed with ubiquitin overexpression. Strains were pulsed with 35S-Met/Cys as described in Figure 2C . Vector: empty vector.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
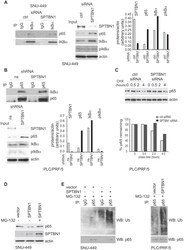
- Figure 3 SPTBN1 induces ubiquitination and degradation of p65. (A and B) Binding of p65 with IkappaBa kept identical in HCC cells upon the inhibition of SPTBN1. SNU-449 cells were transiently transfected with siRNAs as indicated for 48 h (A) and PLC/PRF/5 cells stably transduced with SPTBN1 shRNA or non-specific (ns) shRNA were cultured for 48h (B). Cells were then subjected to immunoprecipitation (IP) and immunoblotting with antibodies as indicated. Whole cell lysates of above cells (inputs) were analyzed by Western blotting by antibodies as indicated. Data are representative of two to three independent experiments with similar results. The intensities of SPTBN1, p65, IkappaBalpha, pIkappaBalpha and actin were measured by ImageJ software, and the relative expression level of each protein/actin was generated. (C) PLC/PRF/5 cells were transiently transfected as indicated for 48h. Cells were then treated with cycloheximide (CHX) for the indicated time periods and were analyzed for the stability of p65 by Western blotting (upper). The intensities of p65 and actin were measured by ImageJ software, and the relative expression units of p65/actin were generated and normalized relative to the expression unit in 0 h. The percentage remaining of p65 expression (compare to zero hour) was then calculated (lower). (D) SNU-449 cells were transfected as indicated for 48 h. Cells were then treated without or with 10 uM MG132 for 4 h before cell lysis and analyzed by Western blotting. (E) SNU
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
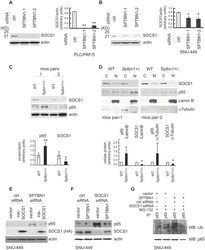
- Figure 4 SPTBN1 enhances the expression of SOCS1, which was required for the regulation of p65 by SPTBN1. (A-B) PLC/PRF/5 (A) or SNU-449 (B) cells were transiently transfected with siRNAs as indicated. Cells were then subjected to Western blot analysis. The relative intensities of SOCS1 to actin in two to three independent Western blotting were analyzed as in Figure 2 A and 2 B. Significance of the difference was evaluated using the Student''s t test (* P < 0.05; ** P < 0.01). (C-D) Liver tissues from two pairs of age- and gender-matched WT and Sptbn1 +/- mice were analyzed by Western blotting (C). Liver tissues from two pairs of age- and gender-matched WT and Sptbn1 +/- mice were lyzed and fractionated into cytoplasmic [C] and nuclear [N] fractions, and subcellular distribution of p65 and SOCS1 in each fraction was assessed by Western blotting (D). The relative intensities of p65 and SOCS1 were analyzed as in Figure 2 A and 2 B. (E) SNU-449 cells were transiently transfected as indicated for 48 h and were then analyzed by Western blot analysis. (F) SNU-449 cells were transiently transfected with control siRNA or siRNA to SOCS1 together with empty vector or SPTBN1 plasmid for 48 h. Cells were then analyzed by Western blot analysis. (G) SNU-449 cells were transiently transfected with control siRNA or siRNA to SOCS1 together with empty vector or SPTBN1 plasmid for 48 h. Cells were then treated with 10uM MG132 for 4 h before cell lysis. Cells were immunoprecipitated with antibod
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
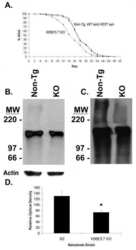
- Fig. 3 PAH content in liver lysates of homozygous and heterozygous Pah-R261Q mice . a Western blots for immunodetection of PAH (alpha-PAH) ( a ) and ubiquitinated protein (alpha-Ub) ( b ) showing the decrease in non-ubiquitinated PAH (non-Ub-PAH; ~51 kDa band) and increase of mono-ubiquitinated PAH (mono-Ub-PAH; ~56 kDa) from genotype Pah WT/WT to Pah R261Q/WT to Pah R261Q/R261Q . The blots are representative from n = 3 replicates for each mice group. GAPDH was used as loading control. c Overview of relative PAH specific activity normalized to activity in Pah WT/WT liver lysates (23.2 +- 2.4 nmol L-Tyr/min/mg protein) ( n = 4 mice for each genotype) and non-Ub-PAH protein (51 kDa) levels from densitometric analysis normalized to both Pah WT/WT liver lysates as well as to GAPDH loading control ( n = 3 mice for each genotype). Data are presented as mean +- SD for Pah WT/WT (purple), Pah R261Q/WT (green), and Pah R261Q/R261Q (ochre), individual values are represented as circles. Differences between genotypes were analyzed by one-way ANOVA followed by Tukey test; differences in PAH activity, p = 0.0005 (***) for Pah WT/WT vs. Pah R261Q/WT , p < 0.0001 (****) for both Pah WT/WT vs. Pah R261Q/R261Q and Pah R261Q/WT vs. Pah R261Q/R261Q ; differences in PAH level, p = 0.0080 (**) for Pah WT/WT vs . Pah R261Q/R261Q . Source data are provided as a Source Data file.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3. UBA6 participates in monoubiquitination of LC3B. ( A ) WT and UBA6-KO H4 cells were incubated with 0, 20 or 50 muM MG132 for 6 hr, and analyzed by SDS-PAGE and immunoblotting with antibodies to LC3B, Ub and beta-tubulin (loading control). ( B ) The ratio of LC3B-I to beta-tubulin was determined from experiments such as that in A. The ratio in WT H4 cells without MG132 treatment was arbitrarily set at 1. Bars represent the mean +- SEM of the ratio from three independent experiments. The indicated p -values were calculated using two-way ANOVA with Tukey's multiple comparisons tests. ( C ) WT and UBA6-KO cells were transfected with plasmids encoding HA-Ub and FLAG-LC3B. Cell lysates were immunoprecipitated (IP) with antibody to the FLAG epitope, and cell lysates and immunoprecipitates were analyzed by SDS-PAGE and immunoblotting with antibodies to the HA and FLAG epitopes. Ubiquitinated FLAG-LC3B can be seen as a faint band on the long exposure of the anti-FLAG blot (arrow). ( D ) H4 cells were transfected with plasmids encoding HA-Ub and FLAG-LC3B. Cells were incubated with 20 muM MG132 for 6 hr before lysis and immunoprecipitation with antibody to the FLAG epitope. Cell lysates and immunoprecipitates were analyzed by SDS-PAGE and immunoblotting with antibodies to the HA and FLAG epitopes. ( E ) Immunofluorescence microscopy showing the localization of FLAG-LC3B and FLAG-LC3B-G120A mutant in WT H4 cells. DAPI (blue) was used to stain the nucleus. Cell edges are outlin
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot