Antibody data
- Antibody Data
- Antigen structure
- References [1]
- Comments [0]
- Validations
- Western blot [1]
- Immunohistochemistry [1]
- Other assay [1]
Submit
Validation data
Reference
Comment
Report error
- Product number
- PA5-57265 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- METTL8 Polyclonal Antibody
- Antibody type
- Polyclonal
- Antigen
- Recombinant full-length protein
- Description
- Immunogen sequence: MNMIWRNSIS CLRLGKVPHR YQSGYHPVAP LGSRILTDPA KVFEHNMWDH MQWSKEEEAA ARKKVKENSA VRVLLEE
- Concentration
- 0.2 mg/mL
Submitted references Methyltransferase METTL8 is required for 3-methylcytosine modification in human mitochondrial tRNAs.
Lentini JM, Bargabos R, Chen C, Fu D
The Journal of biological chemistry 2022 Apr;298(4):101788
The Journal of biological chemistry 2022 Apr;298(4):101788
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
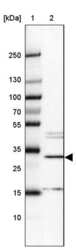
- Experimental details
- Western blot analysis of METTL8 in Lane 1: Marker (kDa) 250, 130, 100, 70, 55, 35, 25, 15, 10; Lane 2: Human cell line HepG2. Samples were probed using a METTL8 Polyclonal Antibody (Product # PA5-57265).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
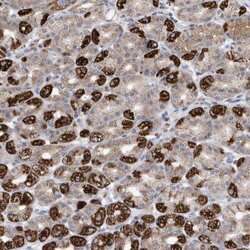
- Experimental details
- Immunohistochemical staining of METTL8 in human stomach tissue shows strong cytoplasmic positivity in parietal cells. Samples were probed using a METTL8 Polyclonal Antibody (Product # PA5-57265).
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
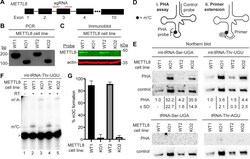
- Experimental details
- Figure 5 METTL8 is required for efficient m 3 C formation in mitochondrial tRNAs in vivo . A , CRISPR/Cas9 gene KO strategy depicting sequence guide RNAs (sgRNAs) targeting exon 3 of the human METTL8 gene. B , genomic PCR demonstrating the loss of exon 3 in METTL8-KO cell lines. Deletion of exon 3 results in a 72 base-pair deletion. C , immunoblot analysis of METTL8 expression in WT versus METTL8-KO human cell lines. Actin was used as a loading control. D , (i) schematic of the Positive Hybridization in the Absence of Modification (PHA) to monitor m 3 C status. PHA probe spans position C32 while the control probe spans a region lacking modifications on the same tRNA. (ii) schematic of primer extension assay to monitor m 3 C modification. E , Northern blot analysis using the PHA assay with probes designed to detect m 3 C at position 32 and a control probe that hybridizes to a different area of the same tRNA. PHA quantification represents the ratio of PHA versus control probe signal expressed relative to the WT1 cell line. Northern blotting and quantification were performed on three independent cell preparations for each cell line with comparable results. F , representative gel from primer extension assays to monitor the presence of m 3 C in mt-tRNA-Thr-UGU from the indicated cell lines. G , quantification of m 3 C formation in ( F ). Primer extension analysis was repeated three times and bars represent the standard deviation from the mean. METTL8, methyltransferase-like 8; mt,
 Explore
Explore Validate
Validate Learn
Learn Western blot
Western blot