Antibody data
- Antibody Data
- Antigen structure
- References [33]
- Comments [0]
- Validations
- Flow cytometry [1]
- Other assay [6]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 11-0458-42 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD45RA Monoclonal Antibody (HI100), FITC, eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: The HI100 monoclonal antibody reacts with human CD45RA, a 220 kDa molecule expressed by subpopulations of CD4+ peripheral T lymphocytes, CD8+ peripheral T lymphocytes, and B cells. The CD45RA+ T cell populations are mainly naive/virgin allowing the use of HI100 mAb as a phenotypic marker to discriminate T cell subsets. Applications Reported: The HI100 antibody has been reported for use in flow cytometric analysis. Applications Tested: This HI100 antibody has been pre-titrated and tested by flow cytometric analysis of normal human peripheral blood cells. This can be used at 5 µL (1 µg) per test. A test is defined as the amount (µg) of antibody that will stain a cell sample in a final volume of 100 µL. Cell number should be determined empirically but can range from 10^5 to 10^8 cells/test. Excitation: 488 nm; Emission: 520 nm; Laser: Blue Laser. Filtration: 0.2 µm post-manufacturing filtered.
- Reactivity
- Human
- Host
- Mouse
- Conjugate
- Green dye
- Isotype
- IgG
- Antibody clone number
- HI100
- Vial size
- 100 Tests
- Concentration
- 5 µL/Test
- Storage
- 4° C, store in dark, DO NOT FREEZE!
Submitted references Fusobacterium nucleatum induces a tumor microenvironment with diminished adaptive immunity against colorectal cancers.
ADAP1 promotes latent HIV-1 reactivation by selectively tuning KRAS-ERK-AP-1 T cell signaling-transcriptional axis.
HLA-independent T cell receptors for targeting tumors with low antigen density.
Dysregulated Peripheral Invariant Natural Killer T Cells in Plaque Psoriasis Patients.
Differences in Maturation Status and Immune Phenotypes of Circulating Helios(+) and Helios(-) Tregs and Their Disrupted Correlations With Monocyte Subsets in Autoantibody-Positive T1D Individuals.
Protective heterologous T cell immunity in COVID-19 induced by the trivalent MMR and Tdap vaccine antigens.
Immune signature of CCR7(+) central memory T cells associates with disease severity and Immunoglobulin E in bronchial asthma.
Heterogeneous disease-propagating stem cells in juvenile myelomonocytic leukemia.
The TCR Repertoire Reconstitution in Multiple Sclerosis: Comparing One-Shot and Continuous Immunosuppressive Therapies.
Single-Cell Analyses Reveal Megakaryocyte-Biased Hematopoiesis in Myelofibrosis and Identify Mutant Clone-Specific Targets.
Serum Amyloid A Proteins Induce Pathogenic Th17 Cells and Promote Inflammatory Disease.
Microglia Require CD4 T Cells to Complete the Fetal-to-Adult Transition.
The downregulated membrane expression of CD18 in CD34(+) cells defines a primitive population of human hematopoietic stem cells.
Frequency determination of breast tumor-reactive CD4 and CD8 T cells in humans: unveiling the antitumor immune response.
MicroRNA‑155 inhibits the proliferation of CD8+ T cells via upregulating regulatory T cells in vitiligo.
Targeting enhancer switching overcomes non-genetic drug resistance in acute myeloid leukaemia.
Effects of Glycogen Synthase Kinase-3β Inhibitor TWS119 on Proliferation and Cytokine Production of TILs From Human Lung Cancer.
TIGIT expressing CD4+T cells represent a tumor-supportive T cell subset in chronic lymphocytic leukemia.
Reciprocal regulation of the Il9 locus by counteracting activities of transcription factors IRF1 and IRF4.
Inherited GINS1 deficiency underlies growth retardation along with neutropenia and NK cell deficiency.
Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection.
The distribution and function of human memory T cell subsets in lung cancer.
Distinct signaling programs control human hematopoietic stem cell survival and proliferation.
Single-cell profiling reveals GPCR heterogeneity and functional patterning during neuroinflammation.
Promoter H3K4 methylation dynamically reinforces activation-induced pathways in human CD4 T cells.
Exhaustion of Activated CD8 T Cells Predicts Disease Progression in Primary HIV-1 Infection.
Activation of Innate and Adaptive Immunity by a Recombinant Human Cytomegalovirus Strain Expressing an NKG2D Ligand.
Alloantigen-specific regulatory T cells generated with a chimeric antigen receptor.
Somatic Variation of T-Cell Receptor Genes Strongly Associate with HLA Class Restriction.
PD-1+Tim-3+ CD8+ T Lymphocytes Display Varied Degrees of Functional Exhaustion in Patients with Regionally Metastatic Differentiated Thyroid Cancer.
Transcriptional profile of tuberculosis antigen-specific T cells reveals novel multifunctional features.
CD55 costimulation induces differentiation of a discrete T regulatory type 1 cell population with a stable phenotype.
Fc receptor-like 3 protein expressed on IL-2 nonresponsive subset of human regulatory T cells.
Kim HS, Kim CG, Kim WK, Kim KA, Yoo J, Min BS, Paik S, Shin SJ, Lee H, Lee K, Kim H, Shin EC, Kim TM, Ahn JB
Frontiers in cellular and infection microbiology 2023;13:1101291
Frontiers in cellular and infection microbiology 2023;13:1101291
ADAP1 promotes latent HIV-1 reactivation by selectively tuning KRAS-ERK-AP-1 T cell signaling-transcriptional axis.
Ramirez NP, Lee J, Zheng Y, Li L, Dennis B, Chen D, Challa A, Planelles V, Westover KD, Alto NM, D'Orso I
Nature communications 2022 Mar 1;13(1):1109
Nature communications 2022 Mar 1;13(1):1109
HLA-independent T cell receptors for targeting tumors with low antigen density.
Mansilla-Soto J, Eyquem J, Haubner S, Hamieh M, Feucht J, Paillon N, Zucchetti AE, Li Z, Sjöstrand M, Lindenbergh PL, Saetersmoen M, Dobrin A, Maurin M, Iyer A, Garcia Angus A, Miele MM, Zhao Z, Giavridis T, van der Stegen SJC, Tamzalit F, Rivière I, Huse M, Hendrickson RC, Hivroz C, Sadelain M
Nature medicine 2022 Feb;28(2):345-352
Nature medicine 2022 Feb;28(2):345-352
Dysregulated Peripheral Invariant Natural Killer T Cells in Plaque Psoriasis Patients.
Hu Y, Chen Y, Chen Z, Zhang X, Guo C, Yu Z, Xu P, Sun L, Zhou X, Gong Y, Yu Q, Shi Y
Frontiers in cell and developmental biology 2021;9:799560
Frontiers in cell and developmental biology 2021;9:799560
Differences in Maturation Status and Immune Phenotypes of Circulating Helios(+) and Helios(-) Tregs and Their Disrupted Correlations With Monocyte Subsets in Autoantibody-Positive T1D Individuals.
Zhang Y, Zhang J, Shi Y, Shen M, Lv H, Chen S, Feng Y, Chen H, Xu X, Yang T, Xu K
Frontiers in immunology 2021;12:628504
Frontiers in immunology 2021;12:628504
Protective heterologous T cell immunity in COVID-19 induced by the trivalent MMR and Tdap vaccine antigens.
Mysore V, Cullere X, Settles ML, Ji X, Kattan MW, Desjardins M, Durbin-Johnson B, Gilboa T, Baden LR, Walt DR, Lichtman AH, Jehi L, Mayadas TN
Med (New York, N.Y.) 2021 Sep 10;2(9):1050-1071.e7
Med (New York, N.Y.) 2021 Sep 10;2(9):1050-1071.e7
Immune signature of CCR7(+) central memory T cells associates with disease severity and Immunoglobulin E in bronchial asthma.
Moaaz M, Youssry S, Baess A, Abed A, Moaaz M
European annals of allergy and clinical immunology 2021 May;53(3):115-127
European annals of allergy and clinical immunology 2021 May;53(3):115-127
Heterogeneous disease-propagating stem cells in juvenile myelomonocytic leukemia.
Louka E, Povinelli B, Rodriguez-Meira A, Buck G, Wen WX, Wang G, Sousos N, Ashley N, Hamblin A, Booth CAG, Roy A, Elliott N, Iskander D, de la Fuente J, Fordham N, O'Byrne S, Inglott S, Norfo R, Salio M, Thongjuea S, Rao A, Roberts I, Mead AJ
The Journal of experimental medicine 2021 Feb 1;218(2)
The Journal of experimental medicine 2021 Feb 1;218(2)
The TCR Repertoire Reconstitution in Multiple Sclerosis: Comparing One-Shot and Continuous Immunosuppressive Therapies.
Amoriello R, Greiff V, Aldinucci A, Bonechi E, Carnasciali A, Peruzzi B, Repice AM, Mariottini A, Saccardi R, Mazzanti B, Massacesi L, Ballerini C
Frontiers in immunology 2020;11:559
Frontiers in immunology 2020;11:559
Single-Cell Analyses Reveal Megakaryocyte-Biased Hematopoiesis in Myelofibrosis and Identify Mutant Clone-Specific Targets.
Psaila B, Wang G, Rodriguez-Meira A, Li R, Heuston EF, Murphy L, Yee D, Hitchcock IS, Sousos N, O'Sullivan J, Anderson S, Senis YA, Weinberg OK, Calicchio ML, NIH Intramural Sequencing Center, Iskander D, Royston D, Milojkovic D, Roberts I, Bodine DM, Thongjuea S, Mead AJ
Molecular cell 2020 May 7;78(3):477-492.e8
Molecular cell 2020 May 7;78(3):477-492.e8
Serum Amyloid A Proteins Induce Pathogenic Th17 Cells and Promote Inflammatory Disease.
Lee JY, Hall JA, Kroehling L, Wu L, Najar T, Nguyen HH, Lin WY, Yeung ST, Silva HM, Li D, Hine A, Loke P, Hudesman D, Martin JC, Kenigsberg E, Merad M, Khanna KM, Littman DR
Cell 2020 Jan 9;180(1):79-91.e16
Cell 2020 Jan 9;180(1):79-91.e16
Microglia Require CD4 T Cells to Complete the Fetal-to-Adult Transition.
Pasciuto E, Burton OT, Roca CP, Lagou V, Rajan WD, Theys T, Mancuso R, Tito RY, Kouser L, Callaerts-Vegh Z, de la Fuente AG, Prezzemolo T, Mascali LG, Brajic A, Whyte CE, Yshii L, Martinez-Muriana A, Naughton M, Young A, Moudra A, Lemaitre P, Poovathingal S, Raes J, De Strooper B, Fitzgerald DC, Dooley J, Liston A
Cell 2020 Aug 6;182(3):625-640.e24
Cell 2020 Aug 6;182(3):625-640.e24
The downregulated membrane expression of CD18 in CD34(+) cells defines a primitive population of human hematopoietic stem cells.
Mesa-Núñez C, Leon-Rico D, Aldea M, Damián C, Sanchez-Baltasar R, Sanchez R, Alberquilla O, Segovia JC, Bueren JA, Almarza E
Stem cell research & therapy 2020 Apr 28;11(1):164
Stem cell research & therapy 2020 Apr 28;11(1):164
Frequency determination of breast tumor-reactive CD4 and CD8 T cells in humans: unveiling the antitumor immune response.
Pinho MP, Patente TA, Flatow EA, Sallusto F, Barbuto JAM
Oncoimmunology 2019;8(8):1607674
Oncoimmunology 2019;8(8):1607674
MicroRNA‑155 inhibits the proliferation of CD8+ T cells via upregulating regulatory T cells in vitiligo.
Lv M, Li Z, Liu J, Lin F, Zhang Q, Li Z, Wang Y, Wang K, Xu Y
Molecular medicine reports 2019 Oct;20(4):3617-3624
Molecular medicine reports 2019 Oct;20(4):3617-3624
Targeting enhancer switching overcomes non-genetic drug resistance in acute myeloid leukaemia.
Bell CC, Fennell KA, Chan YC, Rambow F, Yeung MM, Vassiliadis D, Lara L, Yeh P, Martelotto LG, Rogiers A, Kremer BE, Barbash O, Mohammad HP, Johanson TM, Burr ML, Dhar A, Karpinich N, Tian L, Tyler DS, MacPherson L, Shi J, Pinnawala N, Yew Fong C, Papenfuss AT, Grimmond SM, Dawson SJ, Allan RS, Kruger RG, Vakoc CR, Goode DL, Naik SH, Gilan O, Lam EYN, Marine JC, Prinjha RK, Dawson MA
Nature communications 2019 Jun 20;10(1):2723
Nature communications 2019 Jun 20;10(1):2723
Effects of Glycogen Synthase Kinase-3β Inhibitor TWS119 on Proliferation and Cytokine Production of TILs From Human Lung Cancer.
Tang YY, Sheng SY, Lu CG, Zhang YQ, Zou JY, Lei YY, Gu Y, Hong H
Journal of immunotherapy (Hagerstown, Md. : 1997) 2018 Sep;41(7):319-328
Journal of immunotherapy (Hagerstown, Md. : 1997) 2018 Sep;41(7):319-328
TIGIT expressing CD4+T cells represent a tumor-supportive T cell subset in chronic lymphocytic leukemia.
Catakovic K, Gassner FJ, Ratswohl C, Zaborsky N, Rebhandl S, Schubert M, Steiner M, Gutjahr JC, Pleyer L, Egle A, Hartmann TN, Greil R, Geisberger R
Oncoimmunology 2017;7(1):e1371399
Oncoimmunology 2017;7(1):e1371399
Reciprocal regulation of the Il9 locus by counteracting activities of transcription factors IRF1 and IRF4.
Campos Carrascosa L, Klein M, Kitagawa Y, Lückel C, Marini F, König A, Guralnik A, Raifer H, Hagner-Benes S, Rädler D, Böck A, Kang C, Lohoff M, Garn H, Schaub B, Berberich-Siebelt F, Sakaguchi S, Bopp T, Huber M
Nature communications 2017 May 12;8:15366
Nature communications 2017 May 12;8:15366
Inherited GINS1 deficiency underlies growth retardation along with neutropenia and NK cell deficiency.
Cottineau J, Kottemann MC, Lach FP, Kang YH, Vély F, Deenick EK, Lazarov T, Gineau L, Wang Y, Farina A, Chansel M, Lorenzo L, Piperoglou C, Ma CS, Nitschke P, Belkadi A, Itan Y, Boisson B, Jabot-Hanin F, Picard C, Bustamante J, Eidenschenk C, Boucherit S, Aladjidi N, Lacombe D, Barat P, Qasim W, Hurst JA, Pollard AJ, Uhlig HH, Fieschi C, Michon J, Bermudez VP, Abel L, de Villartay JP, Geissmann F, Tangye SG, Hurwitz J, Vivier E, Casanova JL, Smogorzewska A, Jouanguy E
The Journal of clinical investigation 2017 May 1;127(5):1991-2006
The Journal of clinical investigation 2017 May 1;127(5):1991-2006
Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection.
Eyquem J, Mansilla-Soto J, Giavridis T, van der Stegen SJ, Hamieh M, Cunanan KM, Odak A, Gönen M, Sadelain M
Nature 2017 Mar 2;543(7643):113-117
Nature 2017 Mar 2;543(7643):113-117
The distribution and function of human memory T cell subsets in lung cancer.
Sheng SY, Gu Y, Lu CG, Zou JY, Hong H, Wang R
Immunologic research 2017 Jun;65(3):639-650
Immunologic research 2017 Jun;65(3):639-650
Distinct signaling programs control human hematopoietic stem cell survival and proliferation.
Knapp DJ, Hammond CA, Aghaeepour N, Miller PH, Pellacani D, Beer PA, Sachs K, Qiao W, Wang W, Humphries RK, Sauvageau G, Zandstra PW, Bendall SC, Nolan GP, Hansen C, Eaves CJ
Blood 2017 Jan 19;129(3):307-318
Blood 2017 Jan 19;129(3):307-318
Single-cell profiling reveals GPCR heterogeneity and functional patterning during neuroinflammation.
Tischner D, Grimm M, Kaur H, Staudenraus D, Carvalho J, Looso M, Günther S, Wanke F, Moos S, Siller N, Breuer J, Schwab N, Zipp F, Waisman A, Kurschus FC, Offermanns S, Wettschureck N
JCI insight 2017 Aug 3;2(15)
JCI insight 2017 Aug 3;2(15)
Promoter H3K4 methylation dynamically reinforces activation-induced pathways in human CD4 T cells.
LaMere SA, Thompson RC, Komori HK, Mark A, Salomon DR
Genes and immunity 2016 Jul;17(5):283-97
Genes and immunity 2016 Jul;17(5):283-97
Exhaustion of Activated CD8 T Cells Predicts Disease Progression in Primary HIV-1 Infection.
Hoffmann M, Pantazis N, Martin GE, Hickling S, Hurst J, Meyerowitz J, Willberg CB, Robinson N, Brown H, Fisher M, Kinloch S, Babiker A, Weber J, Nwokolo N, Fox J, Fidler S, Phillips R, Frater J, SPARTAC and CHERUB Investigators
PLoS pathogens 2016 Jul;12(7):e1005661
PLoS pathogens 2016 Jul;12(7):e1005661
Activation of Innate and Adaptive Immunity by a Recombinant Human Cytomegalovirus Strain Expressing an NKG2D Ligand.
Tomić A, Varanasi PR, Golemac M, Malić S, Riese P, Borst EM, Mischak-Weissinger E, Guzmán CA, Krmpotić A, Jonjić S, Messerle M
PLoS pathogens 2016 Dec;12(12):e1006015
PLoS pathogens 2016 Dec;12(12):e1006015
Alloantigen-specific regulatory T cells generated with a chimeric antigen receptor.
MacDonald KG, Hoeppli RE, Huang Q, Gillies J, Luciani DS, Orban PC, Broady R, Levings MK
The Journal of clinical investigation 2016 Apr 1;126(4):1413-24
The Journal of clinical investigation 2016 Apr 1;126(4):1413-24
Somatic Variation of T-Cell Receptor Genes Strongly Associate with HLA Class Restriction.
Klarenbeek PL, Doorenspleet ME, Esveldt RE, van Schaik BD, Lardy N, van Kampen AH, Tak PP, Plenge RM, Baas F, de Bakker PI, de Vries N
PloS one 2015;10(10):e0140815
PloS one 2015;10(10):e0140815
PD-1+Tim-3+ CD8+ T Lymphocytes Display Varied Degrees of Functional Exhaustion in Patients with Regionally Metastatic Differentiated Thyroid Cancer.
Severson JJ, Serracino HS, Mateescu V, Raeburn CD, McIntyre RC Jr, Sams SB, Haugen BR, French JD
Cancer immunology research 2015 Jun;3(6):620-30
Cancer immunology research 2015 Jun;3(6):620-30
Transcriptional profile of tuberculosis antigen-specific T cells reveals novel multifunctional features.
Arlehamn CL, Seumois G, Gerasimova A, Huang C, Fu Z, Yue X, Sette A, Vijayanand P, Peters B
Journal of immunology (Baltimore, Md. : 1950) 2014 Sep 15;193(6):2931-40
Journal of immunology (Baltimore, Md. : 1950) 2014 Sep 15;193(6):2931-40
CD55 costimulation induces differentiation of a discrete T regulatory type 1 cell population with a stable phenotype.
Sutavani RV, Bradley RG, Ramage JM, Jackson AM, Durrant LG, Spendlove I
Journal of immunology (Baltimore, Md. : 1950) 2013 Dec 15;191(12):5895-903
Journal of immunology (Baltimore, Md. : 1950) 2013 Dec 15;191(12):5895-903
Fc receptor-like 3 protein expressed on IL-2 nonresponsive subset of human regulatory T cells.
Nagata S, Ise T, Pastan I
Journal of immunology (Baltimore, Md. : 1950) 2009 Jun 15;182(12):7518-26
Journal of immunology (Baltimore, Md. : 1950) 2009 Jun 15;182(12):7518-26
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Staining of normal human peripheral blood cells with Anti-Human CD45RO PerCP-eFluor® 710 (Product # 46-0457-42) and Mouse IgG2b K Isotype Control FITC (Product # 11-4732-42) (left) or Anti-Human CD45RA FITC (right). Cells in the lymphocyte gate were used for analysis.
- Conjugate
- Green dye
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 1 The distribution of CD4+ and CD8+ T cells subsets in human lung cancer. PBMCs were isolated from the blood of lung cancer patients and healthy donors and analyzed by flow cytometry. a The frequency of the CD3+CD4+ T cells and CD3+CD8+ T cells in the HD-PBMC, PBMCs from healthy donors; NSCLC-PBMC, PBMCs from non-small lung cancer patients, Normal-Ly, from healthy lymph node, NSCLC-Ly, tumor infiltrated lymph node from non-small lung cancer patients. b Representative flow cytometric analyses of CD45RA and CCR7 expression in CD3+CD4+ T cells and CD3+CD8+ T cells, indicating naive T cells (CD45RA+/CD45RO-CCR7+, top right quadrant ), terminal effector T cells (CD45RA+/CD45RO-CCR7-, bottom right quadrant ), central memory T cells (Tcm, CD45RO+/CD45RA-CCR7+, top left quadrant ), and effector memory T cells (Tem, CD45RO+/CD45RA-CCR7-, bottom left quadrant ), gated on the forward and side scatter of the lymphocyte populations. c The frequency and absolute number of the CD4+ ( top ) and CD8+ ( bottom ), Tn ( middle gray ), Teff ( black ), Tcm ( grey ), and Tem ( dark grey ) cell subsets in the blood from the non small cell lung cancer patients and healthy donors. d The events of Tn, Teff, Tcm and Tem cell subsets of CD4+ and CD8+ cells in the blood from non small cell lung cancer patients and healthy donors, expressed as the mean +- SEM. * p < 0.05; ** p < 0.005; *** p < 0.001; Mann-Whitney test (two-tailed) and non-paired Student's t-test
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
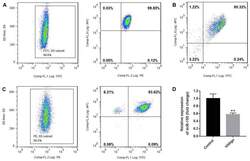
- Figure 1. Purity of CD3 + CD4 + CD45RA + T cells, CD3 + CD8 + T cells and CD4 + CD25 + FoxP3 + Treg cells. CD3 + CD4 + CD45RA + T cells and CD3 + CD8 + T cells were purified by magnetic cell sorting, and their purity was determined by flow cytometry. (A) The purity of CD3 + CD4 + CD45RA + T cells was 99.45% (CD3 + T cells, 99.6%; CD4 + CD45RA + T cells, 99.85%). (B) The purity of CD3 + CD8 + T cells was 95.32%. (C) The purity of CD4 + CD25 + FoxP3 + Treg cells was 93.15% (CD4 + T cells, 99.5%; CD25 + FoxP3 + T cells, 93.62%). (D) miR-155 expression in T cells of the patients with vitiligo and healthy donor was detected by reverse transcription quantitative polymerase chain reaction. **P
- Conjugate
- Green dye
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Fig. 1 Non-genetic adaptation drives clinical resistance in AML. a Meta-analysis from four independent studies analysing either the whole genome or whole exome of AML patients at diagnosis and relapse. Mutations are defined as non-synonymous changes within the coding sequence of any gene. Shared mutations are mutations present at both diagnosis and relapse. Whole exome sequencing data from Li et al. (REF 4 ) was analysed to access the mutations in known AML genes, as defined by the authors. b Schematic of the treatment regime and bone marrow blast percentage for patient BET001 over the clinical trial treatment course (top panel). t-SNE analysis of 7360 individual blast cells isolated from patient BET001 at baseline, remission and relapse (bottom panel). scRNA-seq and genomic DNA sampling points are highlighted on the schematic. c Schematic of treatment regime and bone marrow blast percentage for patient BET002 over the clinical trial treatment course (top panel). t-SNE analysis of 6349 single blast cells isolated from patient BET002 at baseline and relapse (bottom panel). scRNA-seq and genomic DNA sampling points are highlighted on the schematic. d Flow cytometry analysis of cells from patient BET002 at baseline and relapse identifies enrichment for LMPP-like LSCs at relapse based on CD34 + CD38-CD90-CD45RA + expression. Gating strategy is defined by boxes. e Expression analysis of selected LSC signature genes (defined in REF 15 ) in blast cells from patient BET00
- Conjugate
- Green dye
 Explore
Explore Validate
Validate Learn
Learn Flow cytometry
Flow cytometry