14-9908-37
antibody from Invitrogen Antibodies
Targeting: TNFRSF10B
CD262, DR5, KILLER, TRAIL-R2, TRAILR2, TRICK2A, TRICKB
Antibody data
- Antibody Data
- Antigen structure
- References [10]
- Comments [0]
- Validations
- Immunocytochemistry [2]
- Flow cytometry [1]
- Other assay [11]
Submit
Validation data
Reference
Comment
Report error
- Product number
- 14-9908-37 - Provider product page

- Provider
- Invitrogen Antibodies
- Product name
- CD262 (DR5) Monoclonal Antibody (DJR2-4 (7-8)), eBioscience™
- Antibody type
- Monoclonal
- Antigen
- Other
- Description
- Description: The DJR2-4 monoclonal antibody reacts with human DR5, also known as TRAIL-R2, Apo2, TRICK2 and KILLER. DR5 binds to TRAIL, activates NF-kappaB, and induces TRAIL-mediated apoptosis. DR5 is expressed broadly by normal tissues as well as several tumor cells.
- Antibody clone number
- DJR2-4 (7-8)
- Concentration
- 0.5 mg/mL
Submitted references Death agonist antibody against TRAILR2/DR5/TNFRSF10B enhances birinapant anti-tumor activity in HPV-positive head and neck squamous cell carcinomas.
RALB GTPase: a critical regulator of DR5 expression and TRAIL sensitivity in KRAS mutant colorectal cancer.
YIPF2 promotes chemotherapeutic agent-mediated apoptosis via enhancing TNFRSF10B recycling to plasma membrane in non-small cell lung cancer cells.
Targeting the metabolic pathway of human colon cancer overcomes resistance to TRAIL-induced apoptosis.
Combining an Aurora Kinase Inhibitor and a Death Receptor Ligand/Agonist Antibody Triggers Apoptosis in Melanoma Cells and Prevents Tumor Growth in Preclinical Mouse Models.
TRAIL-mediated apoptosis in breast cancer cells cultured as 3D spheroids.
MADD knock-down enhances doxorubicin and TRAIL induced apoptosis in breast cancer cells.
Lack of a functional VHL gene product sensitizes renal cell carcinoma cells to the apoptotic effects of the protein synthesis inhibitor verrucarin A.
A whole-genome RNAi screen identifies an 8q22 gene cluster that inhibits death receptor-mediated apoptosis.
Histone deacetylase inhibitors modulate the sensitivity of tumor necrosis factor-related apoptosis-inducing ligand-resistant bladder tumor cells.
An Y, Jeon J, Sun L, Derakhshan A, Chen J, Carlson S, Cheng H, Silvin C, Yang X, Van Waes C, Chen Z
Scientific reports 2021 Mar 18;11(1):6392
Scientific reports 2021 Mar 18;11(1):6392
RALB GTPase: a critical regulator of DR5 expression and TRAIL sensitivity in KRAS mutant colorectal cancer.
Khawaja H, Campbell A, Roberts JZ, Javadi A, O'Reilly P, McArt D, Allen WL, Majkut J, Rehm M, Bardelli A, Di Nicolantonio F, Scott CJ, Kennedy R, Vitale N, Harrison T, Sansom OJ, Longley DB, Evergren E, Van Schaeybroeck S
Cell death & disease 2020 Oct 29;11(10):930
Cell death & disease 2020 Oct 29;11(10):930
YIPF2 promotes chemotherapeutic agent-mediated apoptosis via enhancing TNFRSF10B recycling to plasma membrane in non-small cell lung cancer cells.
Wang Y, Guo S, Li D, Tang Y, Li L, Su L, Liu X
Cell death & disease 2020 Apr 17;11(4):242
Cell death & disease 2020 Apr 17;11(4):242
Targeting the metabolic pathway of human colon cancer overcomes resistance to TRAIL-induced apoptosis.
Carr RM, Qiao G, Qin J, Jayaraman S, Prabhakar BS, Maker AV
Cell death discovery 2016;2:16067
Cell death discovery 2016;2:16067
Combining an Aurora Kinase Inhibitor and a Death Receptor Ligand/Agonist Antibody Triggers Apoptosis in Melanoma Cells and Prevents Tumor Growth in Preclinical Mouse Models.
Liu Y, Hawkins OE, Vilgelm AE, Pawlikowski JS, Ecsedy JA, Sosman JA, Kelley MC, Richmond A
Clinical cancer research : an official journal of the American Association for Cancer Research 2015 Dec 1;21(23):5338-48
Clinical cancer research : an official journal of the American Association for Cancer Research 2015 Dec 1;21(23):5338-48
TRAIL-mediated apoptosis in breast cancer cells cultured as 3D spheroids.
Chandrasekaran S, Marshall JR, Messing JA, Hsu JW, King MR
PloS one 2014;9(10):e111487
PloS one 2014;9(10):e111487
MADD knock-down enhances doxorubicin and TRAIL induced apoptosis in breast cancer cells.
Turner A, Li LC, Pilli T, Qian L, Wiley EL, Setty S, Christov K, Ganesh L, Maker AV, Li P, Kanteti P, Das Gupta TK, Prabhakar BS
PloS one 2013;8(2):e56817
PloS one 2013;8(2):e56817
Lack of a functional VHL gene product sensitizes renal cell carcinoma cells to the apoptotic effects of the protein synthesis inhibitor verrucarin A.
Woldemichael GM, Turbyville TJ, Vasselli JR, Linehan WM, McMahon JB
Neoplasia (New York, N.Y.) 2012 Aug;14(8):771-7
Neoplasia (New York, N.Y.) 2012 Aug;14(8):771-7
A whole-genome RNAi screen identifies an 8q22 gene cluster that inhibits death receptor-mediated apoptosis.
Dompe N, Rivers CS, Li L, Cordes S, Schwickart M, Punnoose EA, Amler L, Seshagiri S, Tang J, Modrusan Z, Davis DP
Proceedings of the National Academy of Sciences of the United States of America 2011 Oct 25;108(43):E943-51
Proceedings of the National Academy of Sciences of the United States of America 2011 Oct 25;108(43):E943-51
Histone deacetylase inhibitors modulate the sensitivity of tumor necrosis factor-related apoptosis-inducing ligand-resistant bladder tumor cells.
Earel JK Jr, VanOosten RL, Griffith TS
Cancer research 2006 Jan 1;66(1):499-507
Cancer research 2006 Jan 1;66(1):499-507
No comments: Submit comment
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Immunofluorescence analysis of CD262 (DR5) was performed using log phase MDA-MB-231 cells. The cells were fixed with 4% paraformaldehyde for 10 minutes, permeabilized with 0.1% Triton™ X-100 for 10 minutes, and blocked with 1% BSA for 1 hour at room temperature. The cells were labeled with CD262 (DR5) Mouse Monoclonal Antibody (Product # 14-9908-82) at 5 µg/mL in 0.1% BSA and incubated overnight at 4 degree and then labeled with Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor® 488 conjugate (Product # A28175) at a dilution of 1:2000 for 45 minutes at room temperature (Panel a: green). Nuclei (Panel b: blue) were stained with SlowFade® Gold Antifade Mountant with DAPI (Product # S36938). F-actin (Panel c: red) was stained with Rhodamine Phalloidin (Product # R415, 1:300). Panel d represents the merged image showing cytoplasmic localization of CD262 (DR5). Panel e represents control cells with no primary antibody to assess background. The images were captured at 60X magnification.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
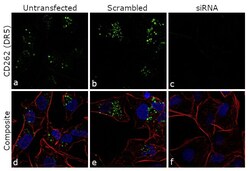
- Knockdown of CD262 (DR5) was achieved by transfecting MDA-MB-231 cells with CD262 (DR5) specific siRNA (Silencer® select Product # s16756). Immunofluorescence analysis was performed using untransfected MDA-MB-231 cells (panels a, d), transfected with non-specific scrambled siRNA (panels b,e) and transfected with CD262 (DR5) specific siRNAs (panel c,f). Cells were fixed, permeabilized, and probed with CD262 (DR5) Monoclonal Antibody (DJR2-4 (7-8)), eBioscience™(Product # 14-9908-82, 5 µg/mL), followed by labelling with Goat anti-Mouse IgG (H+L) Superclonal™ Secondary Antibody, Alexa Fluor 488 (Product # A28175, 1:2000). Nuclei (blue) were stained using ProLong™ Diamond Antifade Mountant with DAPI (Product # P36962) and Rhodamine Phalloidin (Product # R415, 1:300) was used for cytoskeletal F-actin (red) staining. Reduction of specific cytoplasmic localization was observed upon siRNA mediated knockdown (panel c,f) confirming specificity of the antibody to CD262 (DR5). The images were captured at 60X magnification.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Staining of human DR5-transfected cells with 0.5 µg of Mouse IgG1 K Isotype Control Purified (Product # 14-4714-82) (open histogram) or 0.5 µg of Anti-Human CD262 (DR5) Purified (filled histogram) followed by Anti-Mouse IgG Biotin (Product # 13-4013-85) and Streptavidin PE (Product # 12-4317-87). Total viable cells were used for analysis.
Supportive validation
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- NULL
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
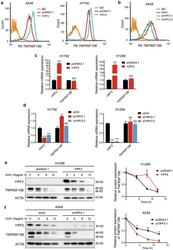
- Fig. 3 YIPF2 enhances TNFRSF10B recycling to plasma membrane. a Overexpression of YIPF2 in A549 and H1792 cells. The surface expression of TNFRSF10B was confirmed by flow cytometry analyses. b Knockdown of YIPF2 expression by YIPF2-1 and YIPF2-2 siRNA in A549 cells. The surface expression of TNFRSF10B was confirmed by flow cytometry analyses. c Relative RT-qPCR analyses of YIPF2 and TNFRSF10B mRNA levels after YIPF2 overexpression in H1792 (left) and H1299 (right) cells ( n = 3). d Relative RT-qPCR analyses of YIPF2 and TNFRSF10B mRNA levels after YIPF2 knocking down in H1792 (left) and H1299 (right) cells ( n = 3). e Left: Overexpression of YIPF2 in H1299 cells in the presence or absence of cycloheximide (CHX) at 10 mug/ml for the indicated times (0, 4, 8 and 12 h). Cell lysates were analyzed by Western blotting with antibodies against YIPF2, TNFRSF10B and ACTB. Right: The band intensity of TNFRSF10B was quantified by ImageJ software and plotted. This experiment was repeated three times independently with similar results. f Left: Knockdown of YIPF2 expression by YIPF2-1 siRNA in A549 cells in the presence or absence of cycloheximide (CHX) at 10 mug/ml for the indicated times (0, 4, 8 and 12 h). Cell lysates were analyzed by Western blotting with antibodies against YIPF2, TNFRSF10B and ACTB. Right: The band intensity of TNFRSF10B was quantified by ImageJ software and plotted. This experiment was repeated three times independently with similar results. (mean +- SEM, n = 3 inde
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
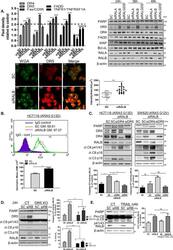
- Fig. 3 RALB silencing regulates DR5 expression levels. A Top Left: HCT116 cells were transfected with 10 nM SC, siRALA, siRALB or siRALGDS for 24 h, following which cells were harvested for protein extraction. Protein was subsequently analysed using a human apoptosis array. Densitometry was performed on the array panels using ImageJ software, and results for DR4, DR5, Fas/CD95, FADD and TNFR1/TNFRSF1A are shown. Top right: PARP, DR5, DR4, FADD, XIAP, Bcl-xL, RALA and RALB levels in CRC cells following RALA, RALB and RALGDS silencing for the indicated times. Bottom left: Wheat germ agglutinin (WGA), a lectin that binds N -acetylglucosamine post-translational modifications on membrane receptors, was used as a plasma membrane marker. Fixed cells were stained with Alexa488-WGA prior to permeabilisation to minimise the staining of glycosylated proteins in the Golgi apparatus. A single confocal image was collected of the Alexa488-WGA stained membrane (green) alongside immunostained endogenous DR5 (red). Laser settings were kept constant between the samples to enable the comparison of membrane-associated DR5 staining in control (SC) and siRALB-treated cells. Bottom right: >20 cells were scored for intensity in ImageJ. The intensity was normalised to cellular area and plotted. The data are representative of three independent experiments. B CRC cells were transfected with 10 nM SC or siRALB for 24 h and DR5 cell-membrane expression was assessed by flow cytometry using a DR5-specific p
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
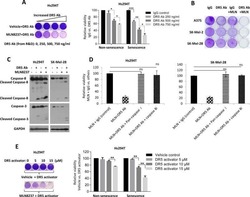
- Figure 3 2DG increases DR5 expression levels with no effect on DR4, however, the increase in TRAIL-induced apoptosis requires additional mechanisms. ( a ) QRT-PCR of DR4 and DR5 transcripts in HT29 cells with and without 2DG treatment. ( b ) Immunoblotting for DR4 and DR5 indicated increased DR5 protein expression with 2DG treatment. ( c ) DR surface expression in various TRAIL resistant colon, gastric and pancreatic adenocarcinoma cells. ( d ) DR5 surface expression increases with 2DG treatment over time in HT-29 cells. ( e ) Apoptosis, as assessed by capsase-3 activation, in the same group of HT-29 cells increased over a 24 h time course upon treatment with 2DG+TRAIL, corresponding temporally with the increase in DR5 expression. ( f ) To knock down DR5 expression, HT-29 cells were transduced with the indicated lentiviral constructs and allowed to grow for 48 h. A lentivirus expressing a scrambled sequence was used as a control. ( g ) Cell death was significantly decreased with DR5 knockdown in 2DG+TRAIL-treated cells, however, remained significantly higher than wild-type cells treated with 2DG or TRAIL alone. n =3, ** indicates P 0.05.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 3 Protein expression of FADD, IAPs, and TRAILR2/DR5 in HPV(+) HNSCC cell lines. ( A ). Protein expression from whole cell lysates of HOK and a panel of HPV(+) HNSCC cell lines were examined by Western blot. Protein expression was quantified by comparison to HOK and actin as a loading control. ( B ) UM-SCC-47 and UPCI-SCC-90 cells were stained with fluorescent conjugated anti-TRAILR2/DR5 antibody and Zombie-violet for viability. The x-axis represents TRAILR2/DR5 fluorescent intensity, and the y-axis represents cell count. Percentages of TRAILR2/DR5 positivity among live cells for each cell line are represented and compared with the negative controls of unstained cells. ( C ) TRAILR2/DR5 staining of Jurkat cells was served as a positive control for data presented by ( B ), and the unstained cells served as the negative control. ( D ) There was no statistical difference in the percentages of TRAILR2/DR5 positive staining cells between the two HNSCC cell lines. The data are presented by six replicates from two independent experiments. * or ** indicates Student''s t test, p < 0.05 or 0.01.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image
- Experimental details
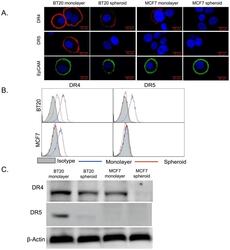
- Figure 1 BT20 and MCF7 spheroids have a lower expression of death receptors. (A) Confocal micrographs of BT20 and MCF7 cells cultured as 3D spheroids and 2D monolayer stained with antibodies against DR4 (red), DR5 (red) and EpCAM (green). (Scale bar = 20 um) (B) Flow cytometry histograms comparing DR4 and DR5 expression in BT20 and MCF7 spheroids and monolayer. (C) Western blot analysis of total DR4 and DR5 expression in cell lysates from BT20 and MCF7 spheroids and monolayer.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 6 COX-2 inhibitor treatment upregulates the expression of DR4 and DR5 in BT20 and MCF7 spheroids. (A) Flow cytometry histograms showing the effect of 100 uM NS-398 treatment on death receptor expression in BT20 and MCF7 cells cultured as spheroids. (B) Western blot analysis of whole cell lysates from untreated and NS-398 treated BT20 and MCF7 spheroids for DR4 and DR5 expression. (C) Flow cytometry scatter plots showing the effect of NS-398 treatment on the prevalence of CD44 hi CD24 lo population in BT20 and MCF7 cells cultured as spheroids.
- Submitted by
- Invitrogen Antibodies (provider)
- Main image

- Experimental details
- Figure 8 COX-2 knockdown reverses TRAIL-resistance in MCF7 spheroid and monolayer cells. (A) qPCR and western blot data showing the relative expression of COX-2 gene (n = 3) and COX-2 protein in control siRNA and COX-2 siRNA transfected MCF7 cells. (B) Flow cytometry histograms and (C) western blot analysis comparing the expression of DR4 in MCF7 cells transfected with control siRNA and COX-2 siRNA cultured as monolayer and spheroids. (D) Bright field images and (E) MTT assay results (n = 5) quantifying the effect of 200 ng/mL of TRAIL in control siRNA and COX-2 siRNA transfected cells cultured as monolayer and spheroids.
 Explore
Explore Validate
Validate Learn
Learn Immunocytochemistry
Immunocytochemistry